| Jan 25, 2021 | |
Characterization of the biomolecular corona at the single nanoparticle level |
|
| (Nanowerk Spotlight) When nanoparticles enter human blood, they come into immediate contact with various biomolecules – proteins, lipids, nucleic acids, metabolites, sugars, etc. These biomolecules form a coating layer on the nanoparticle surface – the so-called biomolecular corona – thereby imparting a unique biological identity to the nanoparticle, which could be very different from the pristine nanoparticle surface (read more in our previous Nanowerk Spotlights: "Exploring the crucial role of biomolecular coronas for nanoparticle-cell interactions" and "Personalized protein coronas result in different therapeutic or toxic impacts of identical nanoparticles"). | |
| The composition and function of the biomolecular corona depends on the physicochemical properties of the nanoparticle, and the composition and incubating conditions of the exposed biological system. | |
| The fact that the properties of a therapeutic nanoparticle can change – to an unknown degree – simply by being introduced into the body, leads to considerable scientific and technological challenges in designing and developing nanomaterials for diagnostic and therapeutic applications. | |
| Consequently, it is essential to understand the interaction of biomolecules in the biomolecular corona with the surface of nanoparticles. | |
| "The concentration and composition of the proteins in the biomolecular corona are traditionally analyzed and quantified by liquid chromatography-tandem mass spectroscopy (LC-MS/MS) and proteomics," Morteza Mahmoudi, an Assistant Professor in the Precision Health Program at Michigan State University, tells Nanowerk. "Although these techniques provide reliable data on the function and structure of the proteins associated with the biomolecular corona, they cannot show the distribution and interaction of proteins and other macromolecules within the biomolecular corona." | |
| In a new study, reported in Nature Communications ("Nanoscale characterization of the biomolecular corona by cryo-electron microscopy, cryo-electron tomography, and image simulation"), Mahmoudi and his collaborators used high-resolution transmission electron microscopy (HRTEM) of air-dried negative-stained model nanoparticles (i.e., carboxylated polystyrene), cryo-electron microscopy (cryo-EM) and cryo-electron tomography (cryo-ET) of the nanoparticles in their native state, three-dimensional (3D) reconstruction, and image analysis to visualize macromolecules and their distribution and geometric arrangement on the surface of individual nanoparticles. | |
 |
|
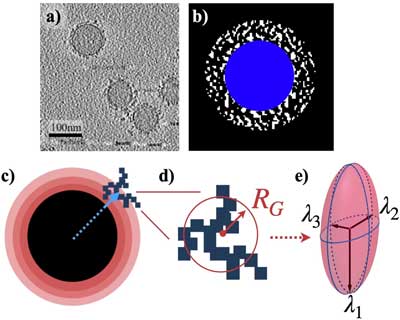
| Defining the physical characteristics of biomolecular corona. a) A tomographic slice shows the biomolecular corona (BC) surrounding the nanoparticles (NPs). The large circles represent spherical NPs. The BC is represented by black points forming clusters around the NPs. The horizontal black bar-like shapes in the bottom right corner of the image are artifacts of smaller gold NPs (used for focusing purposes) that are used to measure the scale of the image. b) A binarized image showing the BC (white) surrounding the NP (blue). c) The space around the NP is divided into spherical shells. The blue arrow points to the center of the cluster. The position of each cluster was determined according to where its center is located among the various shells. d) A schematic representation of an arbitrary cluster. The size of the cluster was characterized by a sphere with a radius of gyration RG. Shown here is a 2D representation. e) The eigenvalues of the gyration tensor are represented with the radii of a hypothetical ellipsoid that best fits the cluster. (Reprinted under a Creative Commons Attribution 4.0 International License) | |
| "The uniqueness of our findings is the morphological detail of the biomolecules, their distribution, and association with the surface of our model polystyrene nanoparticles at the nanometer scale," Mahmoudi reports. "Although the characteristics of the biomolecular corona within the same sample appear homogeneous (i.e., dimensions of molecular clusters, anisotropy, asphericity distribution, and eigenvalues of the gyration tensor), significant variations exist in the distribution and association of the macromolecules among individual nanoparticles. This may significantly affect the biological fate of the nanoparticles." | |
| The team's results also reveal the presence of biomolecular residues between and around the nanoparticles that are not associated with the biomolecular corona. | |
| Since the sample preparation procedure for the LC-MS/MS analysis was applied to cryo-EM and cryo-ET, the researchers think it is likely the data obtained by the LC-MS/MS is an average of all the proteins in the corona of the nanoparticles and the nonspecific proteins remaining in the residue of the plasma. | |
| This study enables more accurate analysis and interpretation of the biomolecular corona at the level of a single nanoparticle when compared to conventional quantitative results obtained from gel electrophoresis and LC-MS/MS. | |
| These latter techniques represent averages of biomolecular corona formed on all nanoparticles, which may not sufficiently define the biological identity of individual nanoparticles. | |
| The 3D reconstructed Cryo Electron Tomography volumes of the reconstructed biomolecular corona structure – blue and yellow colors show nanoparticles and protein clusters, respectively. (Video: Mahmoudi Lab, Michigan State University) | |
| The analysis of biomolecular corona at the level of a single nanoparticle and its variability across nanoparticles in the same sample, as well as the discovery of the presence of nonspecific proteins in the plasma residue enable better characterization of the nanoparticles and improve predictions of safety and biological efficacy of nanoparticles. | |
| "Last but not least, our findings also suggest that the employed combination of the electron microscopy techniques and image analysis can be used as a gold standard for defining 1) the purity, and 2) homogeneity/heterogeneity of the biomolecular corona at nanoscale resolution," Mahmoudi concludes. "The former helps nanomedicine community to define the accuracy and reliability of proteomics and analytical chemistry data of the biomolecular corona at the surface of nanoparticles; the latter is important for defining the suitability of various types of nanoparticles for clinical applications. This means that nanoparticles with a high degree of heterogeneity should not be used in human applications as those individual nanoparticles across the same sample will have significantly different biological fate, pharmacokinetic, and toxicity." | |
 By
Michael
Berger
– Michael is author of four books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology (2009),
Nanotechnology: The Future is Tiny (2016),
Nanoengineering: The Skills and Tools Making Technology Invisible (2019), and
Waste not! How Nanotechnologies Can Increase Efficiencies Throughout Society (2025)
Copyright ©
Nanowerk LLC
By
Michael
Berger
– Michael is author of four books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology (2009),
Nanotechnology: The Future is Tiny (2016),
Nanoengineering: The Skills and Tools Making Technology Invisible (2019), and
Waste not! How Nanotechnologies Can Increase Efficiencies Throughout Society (2025)
Copyright ©
Nanowerk LLC
|
|

Become a Spotlight guest author! Join our large and growing group of guest contributors. Have you just published a scientific paper or have other exciting developments to share with the nanotechnology community? Here is how to publish on nanowerk.com. |
|


