| Posted: Feb 06, 2015 |
Diamond microscope
|
|
(Nanowerk News) Proteins are the building blocks of all living things, and they exist in virtually unlimited varieties, most of whose highly complex structures have not yet been determined. Those structures could be key to developing new drugs or to understanding basic biological processes.
|
|
But figuring out the arrangement of atoms in these complicated, folded molecules usually requires getting them to form crystals large enough to be observed in detail — and for many proteins, that is either impossible or dauntingly difficult.
|
|
Now a new technique being developed by researchers at MIT and elsewhere shows great promise for producing highly detailed images of individual proteins, no matter how complicated their structure, without the need for crystallization. The findings are described in the journal Physical Review X ("Atomic-Scale Nuclear Spin Imaging Using Quantum-Assisted Sensors in Diamond") by MIT graduate student Ashok Ajoy, postdoc Ulf Bissbort, associate professor of nuclear science and engineering Paola Cappellaro, and others at MIT, the Singapore University of Technology and Design, and Harvard University.
|
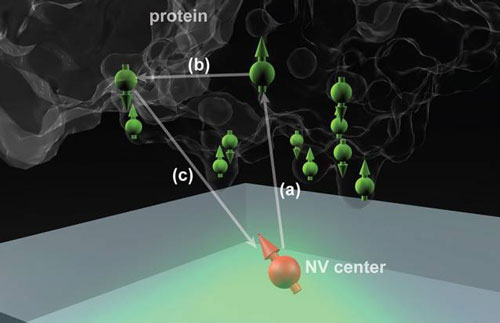 |
| Nitrogen vacancy (NV) centers in diamond could potentially determine the structure of single protein molecules at room temperature. Here the NV center is 2 to 3 nanometers below the surface, and the protein molecule is placed above it. (Image courtesy of the researchers)
|
|
The technique makes use of microscopic defects within the crystal structure of diamond — defects that can be induced, in a controlled way, in the lab. These defects, called nitrogen-vacancy (NV) centers, occur when nitrogen atoms are introduced into the crystal structure, each replacing one carbon atom in a perfectly spaced diamond lattice.
|
|
Such lattices may also include naturally occurring vacancies — imperfections where a carbon atom is missing from its normal place in the lattice. When a nitrogen atom and a vacancy come together, they form an NV center, which can be harnessed to detect the position and attributes — specifically, the spin states — of protons and electrons in atoms placed very close to them.
|
|
That’s done by shining laser light at the diamond surface, which causes the NV centers to fluoresce. By detecting and analyzing the emitted light, it is possible to reconstruct details of the spin of nearby particles.
|
|
The ability to use NV centers in diamond has developed in the last few years, Ajoy says, and many groups are now working to make use of them for applications in quantum computation and quantum communication. When the NV centers are very close to a diamond’s surface — within a few nanometers — they can also be used to sense the spin states of particles within a molecule placed on the surface. The individual atoms and their positions can then, in principle, be detected and mapped out, revealing the molecular structure.
|
|
The idea is to “place a biological molecule on top of the diamond, and try to determine its structure,” Ajoy explains. With proteins, “the structure and function are closely related,” he says, so being able to map out that structure precisely could help in understanding both how some basic biological processes work, and how new drugs might be developed to interact with specific molecular targets.
|
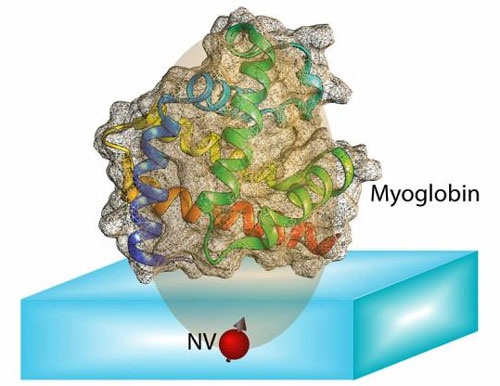 |
| The spin of an NV center can be completely polarized optically. The transfer of this polarization from the NV center to nuclear spins in the protein molecule allows us to unravel couplings between spins in the molecule. Protein structure can be computed from information contained in these couplings. (Image courtesy of the researchers)
|
|
“It could help in developing something that fits on or around [a target molecule], or blocks it,” Ajoy says. “The first step is to know the structure.”
|
|
Efforts to decode the molecular structure of proteins have mostly used X-ray crystallography, transmission electron microscopy, or nuclear magnetic resonance. But all of these methods require large sample volumes — for example, X-ray diffraction requires aggregating the molecules as crystals — so none of them can be used to study individual molecules. This greatly limits the applicability of such techniques.
|
|
“There are many molecules where this doesn’t work out, because you can’t grow the crystals, or they are very hard to grow,” Ajoy says. “For these molecules, our method might be useful because you don’t need the crystals, you just need a single molecule.”
|
|
What’s more, while other techniques require specialized conditions such as very low temperatures or a vacuum, the new technique “perhaps can determine structure at room temperature, under ambient conditions,” he says.
|
|
The work so far is theoretical; the next step, which the team has already begun, is to produce actual images based on this technique. “We started building this setup a year ago, and we have preliminary experiments,” Ajoy says. Actual images of molecules are probably still a few years away, he says.
|
|
This technique is a “very important breakthrough in the development of new techniques for biomolecular structure determination,” says Fedor Jelezko, a professor of quantum optics at the University of Ulm, in Germany, who was not involved in this research. “This is excellent work which will provide very high impact on many fields of science. I am sure that several groups around the world will attempt to realize experiments” based on the ideas in this paper.
|


