| Jun 03, 2016 |
FTIR and microarrays: Enabling more information from less sample
|
|
(Nanowerk News) By using Fourier Transform Infrared Spectroscopy (FTIR), researchers at the Center for Structural Biology and Bioinformatics, Université Libre de Bruxelles, Brussels, Belgium can greatly increase the amount of information that can be extracted from a protein microarray. In a new report in the current issue of Biomedical Spectroscopy and Imaging ("Analysis of protein microarrays by FTIR imaging"), they show how high-quality spectra can be obtained from spots of protein no larger than the diameter of a human hair.
|
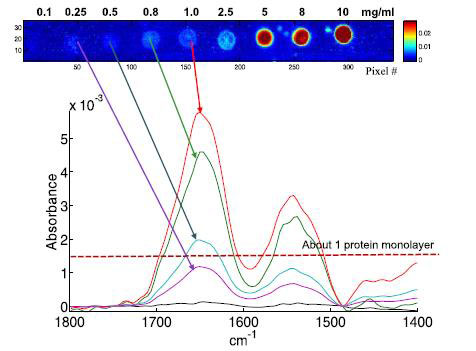 |
| Example of single pixel spectra extracted from albumin spots obtained from 1.0, 0.8, 0.5 and 0.25 mg/ml solutions, as indicated by the arrows. The concentration resulting, on the average, in a 1 protein monolayer is indicated by the dotted line. (© Biomedical Spectroscopy and Imaging)
|
|
The common use of protein microarrays requires the binding of proteins to other compounds such as therapeutic drugs. A fluorescent molecule is attached to the protein so that if binding with the drug occurs, there will be a light signal from the bound pair. However, these fluorescent proteins can be difficult and expensive to produce, and the information is limited to determining whether the drug binds or not.
|
|
Infrared (IR) spectroscopy can probe the molecular structure of a substance. Different wavelengths of infrared light are absorbed by different chemical bonds in a molecule, and by scanning a range of wavelengths, the kinds of bonds can be measured. These measurements comprise a fingerprint of the molecules in the sample. Furthermore, IR spectra account not only for the chemical nature of cell molecules, but also for their shape. They are particularly very sensitive to protein secondary structure.
|
|
Researchers created microarrays using a commercial tool in which about 100 picoliters of protein were deposited from solution. The resulting spots of protein were about 100 micrometers in diameter. A 128x128 focal-plane-array was used to collect a full infrared spectrum from each of the spots in the array, resulting in 16,384 complete spectra. These spectra were preprocessed to remove random noise and do background correction.
|
|
In order to test the sensitivity of the method, lysozyme, albumin, and hemoglobin solutions were prepared at concentrations from 10 to 0.1 mg/ml and deposited in multiple spots. Single spots provided structural and concentration information from a typical protein, albumin.
|
|
According to lead investigator Erik Goormaghtigh, PhD, the combination of FTIR and microarray technology has three main advantages.
|
|
1. Label-free detection: Labelled proteins may be not commercially available or very expensive. It also enhances quality because the labeling procedure often destroys part of the protein structure.
|
|
2. Direct and absolute quantification of proteins: Infrared detection can quantify the amount of bound protein in a sample.
|
|
3. Full imprint of proteins: Infrared imaging provides a complete vibrational spectrum of the binding molecule, which includes information on chemical reactions and protein secondary structure.
|
|
"The results of this study show that high quality spectra can be obtained from minute amounts of proteins, i.e. below a single monolayer of proteins," explained Dr. Goormaghtigh. "This is important as it opens the way to use infrared imaging, instead of fluorescence for instance, for detection of binding. This work also shows that it is now possible to produce high throughput protein analysis by combination of microarrays technology and infrared spectroscopy imaging allowing hundreds of proteins to be quantitatively analyzed in a few minutes. The whole process from protein production/purification to microarray analysis is now integrated in a high throughput robotic platform called Robotein."
|
|
"The study by Goormagtigh and co-workers will further enhance the usefulness of FTIR spectroscopy for protein pharmaceuticals and proteomics research. The approach will nicely complement and supplement existing techniques, used for protein characterization, such as CD, NMR and mass spectrometry," commented Biomedical Spectroscopy and Imaging Editor-in-Chief Dr. Parvez I. Haris, Faculty of Health & Life Sciences, De Montfort University, UK.
|

