| Posted: Jul 24, 2013 | |
Optical detection of epigenetic marks |
|
| (Nanowerk Spotlight) Understanding the purpose of the molecular modifiers that annotate DNA strands – called epigenetic markers – and how they change over time will be crucial in understanding biological processes ranging from embryo development to aging and disease. But just how the markers work, and what different markers mean, is painstaking work that still has left a long way to go. | |
| Deciphering the epigenetic code is a massive mapping exercise, but will provide important information on how epigenetic markers differ among cell types and between healthy and sick individuals. Actually detecting the markers, however, has proved difficult. | |
| Advancing this research field, scientists have now reported the first direct visualization of individual epigenetic modifications in the genome. This is a technical and conceptual breakthrough as it allows not only to quantify the amount of modified bases but also to pin point and map their position in the genome. | |
| "Being able to count individual modifications results in the most sensitive detection of genomic hydroxymethylcytosine ever reported," Yuval Ebenstein, who leads the NanoBioPhotonics research group at Tel Aviv University, tells Nanowerk. "Our findings open up research into numerous human cell types,including human blood cells, that so far have not been accessible due to low levels of epigenetic modification. | |
 |
|
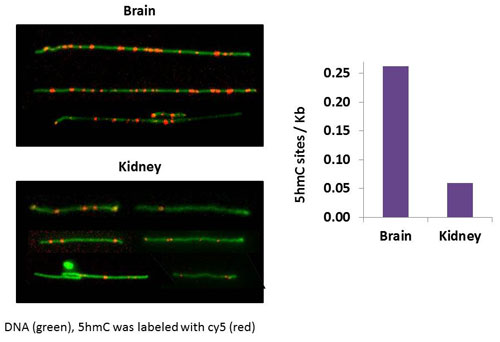
| Single molecule quantification of 5hmC in genomic DNA extracted from mouse tissue. (Image: Ebenstein Lab, Tel Aviv University) | |
| Ebenstein's team reported their findings in Chemical Communications ("Optical detection of epigenetic marks: Sensitive quantification and direct imaging of individual hydroxymethylcytosine bases"). | |
| In this paper, the scientists demonstrate the utility of optical sensing for the detection of and quantification of 5-hydroxymethylcytosine (5hmC) residues in genomic DNA. | |
| Since its discovery in human genomic DNA in 2009, this modification has been the focus of numerous studies and has been established as an important epigenetic mark linked to stem-cell differentiation, aging, and neurodegenerative disease. | |
| "In our work, we utilize enzymatic glucosylation followed by click labeling of a fluorescent reporter to specifically tag individual 5hmC sites along DNA molecules," Ebenstein explains. "The result is a simple yet highly sensitive 5hmC quantification assay that is compatible with bulk DNA analysis as well as with 5hmC detection on individual DNA molecules. | |
| With this report, the group extends their previous work on mapping DNA binding proteins to mapping epigenetic DNA modifications, both of which we covered here in our Nanowerk Spotlight series: "DNA reference tags allow single-molecule research on complex genomes" and "Quantum dots light up individual DNA binding proteins". | |
 |
|
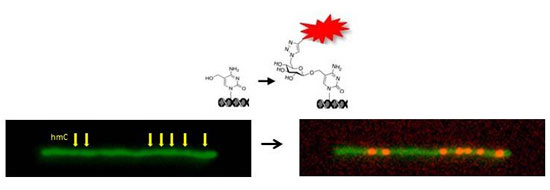
| An engineered 5hmC pattern on lambda phage DNA is visualized after enzymatic glucosylation and fluorescent labeling. (Image: Ebenstein Lab, Tel Aviv University) | |
| The immediate impact of these results is the demonstration that a simple and quick UV-vis measurement can replace tedious radioactive, mass spectrograph and affinity based methods existing today – basically enabling any lab equipped with a UV-vis spectrophotometer to quantify hydroxymethylation levels. | |
| "The more exciting prospect is to be able to profile individual genomic molecules and detect distinct hmC patterns that may serve as biomarkers for disease," says Ebenstein. | |
| "Optical mapping of DNA has mainly focused on genetic analysis and tremendous progress has been made in recent years," he adds. "Our work enables the addition of an epigenetic overlay to existing fluorescence barcoding technologies in order to map epigenetic patterns in the context of the underlying sequence. The main challenge now is to make sense of the information and new analysis algorithms must be developed." | |
 By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
|
|
|
Become a Spotlight guest author! Join our large and growing group of guest contributors. Have you just published a scientific paper or have other exciting developments to share with the nanotechnology community? Here is how to publish on nanowerk.com. |
|
