| Posted: Feb 20, 2018 | |
The ongoing development of DNA nanotechnology |
|
| (Nanowerk Spotlight) DNA, the fundamental building block of our genetic makeup, has become an intense nanotechnology research field. DNA nanotechnology is based on the idea that DNA can be used as a programmable structural material. | |
| Nucleotide interaction is highly specific following the rule of Watson–Crick base pairing (i.e., A-T and G-C paring), which allows rational placement and assembly of DNA strands relative to each other. DNA strands also contain chemically tunable groups, which make it easy to functionalize DNA with molecules and nanomaterials. As a result, DNA nanostructures can encode both structural and chemical/functional information. More importantly, for the purpose of surface patterning, the cost of DNA nanostructure is not a major concern. | |
| Nanotechnology researchers use it to create artificial rationally designed nanostructures for diverse applications in biology, chemistry, and physics. DNA could be particularly useful for the design of electric circuits and DNA-based computing because it self-assembles, it self-replicates and it can adopt various states and conformations (read more: "DNA electronics in nanotechnology"). | |
| Compared to the traditional long-term storage method that uses pizza-sized reels of magnetic tape, DNA storage is potentially less expensive, far more physically compact, more energy efficient, and longer lasting—DNA survives for hundreds of years and doesn’t require maintenance. Files stored in DNA also can be very easily copied for negligible cost. | |
| DNA’s storage density is staggering (read more: Translation software enables efficient DNA data storage). Consider this: humanity will generate an estimated 33 zettabytes of data by 2025 – that’s 3.3 followed by 22 zeroes. All that information would fit into a ping pong ball, with room to spare. The Library of Congress has about 74 terabytes, or 74 million million bytes, of information – 6000 such libraries would fit in a DNA archive the size of a poppy seed. Facebook’s 300 petabytes (300,000 terabytes) could be stored in half a poppy seed. | |
| Then of course there is a rich body of work on DNA use in nanorobotics ("Nanorobotic arm to operate within DNA sequence") and nanofabrication such as Ned Seeman's work at NYU (see for instance: "Researchers send DNA on sequential, and consequential, building mission" or "Scientists use DNA origami to create 2D structures" or "RNA used to control a DNA rotary nanomachine"). | |
| At present, an ever-increasing number of research groups are exploiting programmable self-assembly properties of nucleic acids in creating rationally designed nanoshapes and nanomachines for many different uses. | |
| In a progress report in Advanced Materials ("Evolution of Structural DNA Nanotechnology"), scientists from Aalto University in Finland focus on the different design paradigms (DNA origami and the related techniques are particularly emphasized), selected high-quality shapes, and the software that enable user-friendly design and fabrication of DNA nanoobjects. | |
| One of the most important developments in structural DNA nanotechnology has been the use of a scaffold strand and hundreds of short staple strands for the assembly of three-dimensional (3D) DNA origami objects. These objects are similar in size to virus capsids and have gained a lot of interest as nanocontainers for drug delivery. Since they can be programed to carry out specific tasks (like transporting molecules and releasing them at a target site), these assemblies act like a robotic system – nanorobots. | |
| The review paper features an extensive discussion on the origin of DNA origami, its evolution from 2D to 3D, and the trend towards building larger structures beyond the nanoscale. | |
 |
|
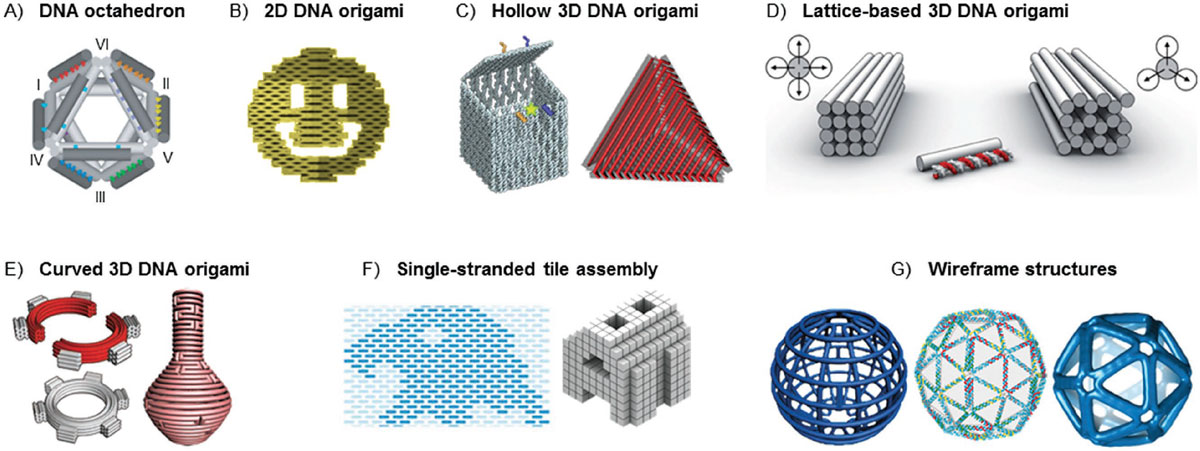
| DNA origami and beyond. A) DNA octahedron that inspired development of DNA origami. B) 2D DNA origami (smiley face serves as an example). C) Hollow 3D DNA origami shapes that are folded from 2D origami sheets. D) Lattice-based 3D origami (square and honeycomb lattice). E) 3D origami with twists and bends. F) Single-stranded tile-based assembly in 2D and in 3D. G) Wireframe-based DNA structures. (© Wiley-VCH Verlag) (click on image to enlarge) | |
| Another big section of the review is dedicated to design software and simulation of the DNA nanoshapes. | |
| The first design software for DNA nanostructures was written by Ned Seeman in 1985 (Journal of Molecular Graphics, "Interactive design and manipulation of macro-molecular architecture utilizing nucleic acid junctions") and succeeded by a better algorithm he developed in 1990 (Journal of Biomolecular Structure and Dynamics, "De novo design of sequences for nucleic acid structural engineering"). | |
| In 2006, Ned Seeman and colleagues developed GIDEON, which was later on partially replaced with other software and techniques – however, GIDEON is still used in many labs. After the dawn of DNA design software, the researchers quickly adopted new and revolutionary DNA origami method, and subsequently, this effort yielded first origami-oriented software SARSE and soon after even more advanced tools. | |
| caDNAno by Douglas et al. and CanDo by Castro et al. and Kim et al. are widely used together for designing 2D and 3D DNA origami nanostructures and computationally predicting the actual shapes of the designed structures in aqueous solution. The core purpose of caDNAno is to simplify and speed up the design process of DNA nanostructures. CanDo is a tool for predicting appearance, mechanical fluctuations, and flexibility, as well as twists and bends of the particular shape in the solution. The software was developed by Hendrik Dietz's (Technische Universität München) and Mark Bathe's groups (MIT) and it is currently maintained by the Laboratory for Computational Biology and Biophysics at MIT. | |
| The authors also discuss additional programs Tiamat, vHelix and DAEDALUS. | |
| With regard to practical applications, the researchers caution that, despite the rapid progress in designing versatile structures and in developing fabrication methods reported in their paper, many implementations of DNA nanotechnology are still quite impractical. | |
| However, there already exist implementations for helping researchers to map the atomic structure of proteins, computing in living cells, and soon, prospectively, tracking and curing diseases with DNA structures might become possible. | |
| In the review's final section, the authors summarize the latest advances in the design optimization and list some recent and notable examples how DNA nanotechnology can be harnessed in a number of subfields of materials science, biotechnology, and medicine. | |
| In particular, they remark on the combination of top-down with bottom-up nanofabrication to fabricate biosensors, nanopores, optics, plasmonics, and molecular electronics; molecular computing, signaling and robotics; and finally nanobiomedicine and characterization of pivotal biomolecules. | |
| "The new designs can become key players in rapid progress of various other research fields diverging from materials science to nanobiomedicine," the authors conclude their review. "Currently nearly 300 laboratories are working in the field, but as the design methods are becoming more and more accessible for users with different backgrounds, the number of participating groups is ever increasing. Therefore, this development will be highly important for creating novel biomaterials and cutting-edge applications." | |
 By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
|
|
|
Become a Spotlight guest author! Join our large and growing group of guest contributors. Have you just published a scientific paper or have other exciting developments to share with the nanotechnology community? Here is how to publish on nanowerk.com. |
|
