| Nov 03, 2022 | |
Incongruent results from proteomics core facilities compromise nanoparticle protein corona research |
|
| (Nanowerk Spotlight) The protein corona plays a critical role not only in clinical translation of nanomaterials but also in a wide range of other nanomedicine areas such as cellular uptake, targeting, drug delivery, and theranostics applications. | |
| A protein corona develops when nanoparticles enter human blood and come into contact with various biomolecules such as proteins, lipids, nucleic acids, metabolites, and sugars. The coating layer on the nanoparticle surface that these biomolecules form is called the biomolecular protein corona. | |
| Its significance for biomedical applications, for instance nanomedicine drug delivery, lies in its role of imparting a unique biological identity to the nanoparticle, which could be very different from the pristine nanoparticle surface. | |
| The complex nature of the protein corona composition significantly affects the biodistribution, targeting efficacy and also toxicity as protein corona gives a new biological identity to nanoparticles. In other word, protein corona is the 'visible side' of nanoparticles and therefore, has a significant impact on the in vivo environment. | |
| The fact that the properties of a therapeutic nanoparticle can change – to an unknown degree – simply by being introduced into the body, leads to considerable scientific and technological challenges in designing and developing nanomaterials for diagnostic and therapeutic applications. | |
| This means that it is imperative for the diagnostic and therapeutic safety and efficacy of nanomedicines to understand the interaction of biomolecules in the biomolecular corona with the surface of nanoparticles. | |
| "Robust characterization of the identity and abundance of the protein corona is entirely dependent on liquid chromatography coupled to mass spectroscopy (LC-MS/MS)," Morteza Mahmoudi, an Assistant Professor in the Precision Health Program at Michigan State University, tells Nanowerk. "Unfortunately, the variability of this technique for the purpose of protein corona characterization remains poorly understood." | |
| To improve this situation, Mahmoudi and his team investigated the variability of LC-MS workflows in protein corona results by sending 17 identical aliquots of protein coronas to different proteomics core facilities across the US and analyzed the retrieved datasets. | |
| They found that, while the shared data between the cores correlate well, there is a significant heterogeneity in the data retrieved from different cores. Surprisingly, out of 4022 identified unique proteins, only 73 (1.8%) were shared across different core facilities. | |
| The results, reported in Nature Communications ("Measurements of heterogeneity in proteomics analysis of nanoparticle protein corona across core facilities"), highlight the critical effect of LC-MS details – e.g., sample preparation protocol, instrumentation, and raw data processing – on protein corona results, which can create bias in interpretation of protein corona applications including biomarker discovery and biological fate of nanomedicine technologies. | |
| These findings reveal that protein corona datasets cannot be easily compared across independent studies and more broadly compromise the interpretation of protein corona research, with implications in biomarker discovery as well as the safety and efficacy of nanoscale biotechnologies. | |
 |
|
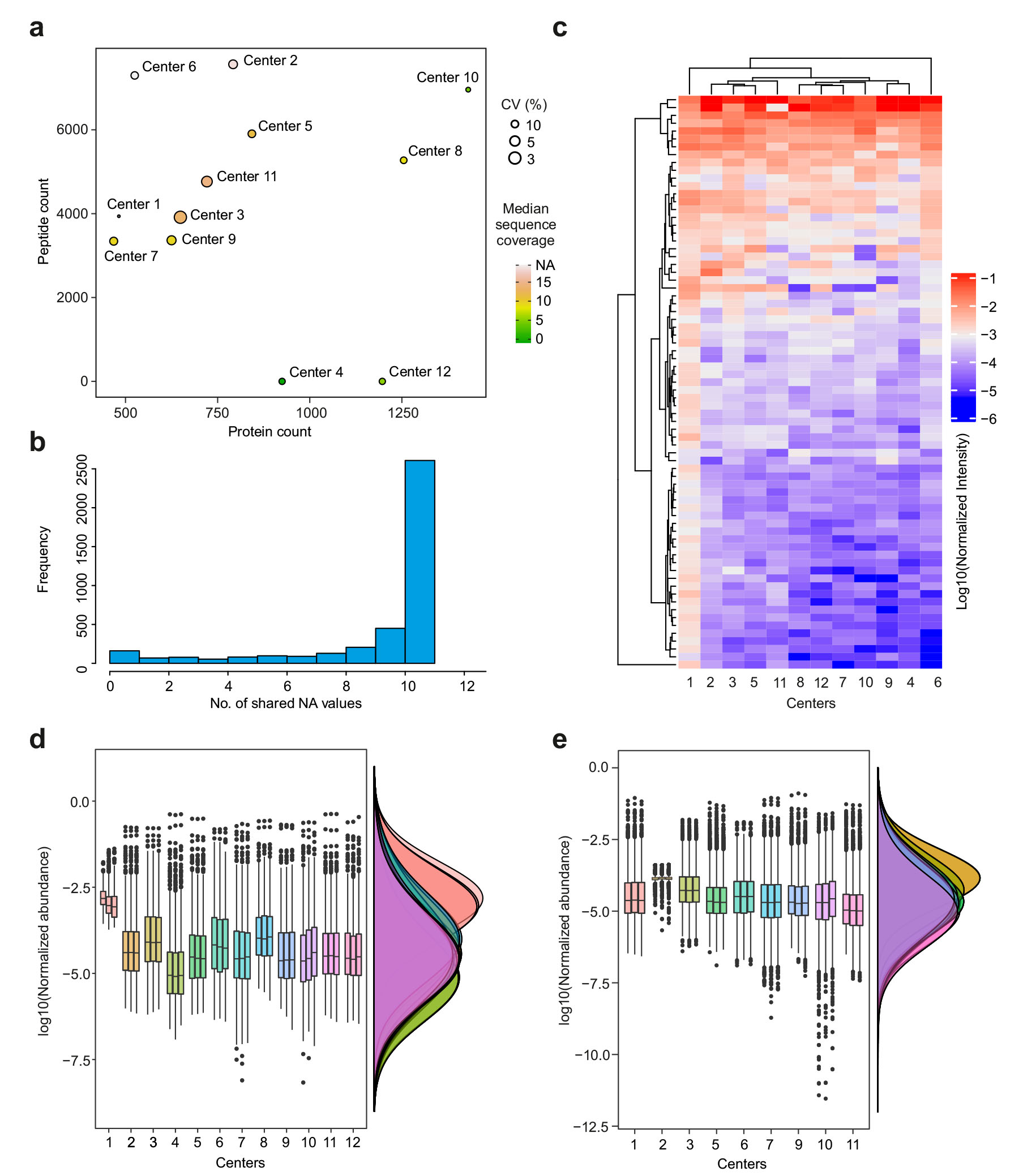
| Comparative protein corona proteomics analysis by the 12 cores that provided semiquantitative protein intensity information. a Comparing the average peptide count (peptide count was not provided by cores 4 or 12), average protein count, median coefficients of variation CV (%) between the technical replicates and median protein sequence coverage (%) for 12 core facilities. b The number of missing values expectedly increased when data from different cores were merged. c Heatmap of 73 shared proteins with any number of peptides between the 12 cores. d Distribution of protein-level intensities for the 12 cores. e Distribution of peptide level intensities for 10 cores, as cores 4 and 12 did not provide peptide level intensities (center line,median; box limits contain 50%; upper and lower quartiles, 75 and 25%; maximum, greatest value excluding outliers; minimum, least value excluding outliers; outliers,more than 1.5 times of upper and lower quartiles). All analyses were based on three technical replicates. (CC BY-NC-ND, Nature Communications) | |
| Mahmoudi's team already reported several overlooked factors at the nanobio interfaces and composition of protein corona such as number and type of the proteins), which is one of the critical issues in prediction of the safety, biodistribution, and diagnostic/therapeutic efficacy of nanomedicine products (read more in our previous Nanowerk Spotlights: "Exploring the crucial role of biomolecular coronas for nanoparticle-cell interactions"; "Personalized protein coronas result in different therapeutic or toxic impacts of identical nanoparticles" and "Characterization of the biomolecular corona at the single nanoparticle level"). | |
| "Currently, the characterization of protein corona, in terms of the types and abundance of proteins, totally depends on a single technique (i.e., LC-MS), and the protein corona results and the corresponding interpretations are significantly different when using different LC-MS instrumentation," Mahmoudi says. "What we found in our study is that there is a remarkable variation in the protein corona composition of similar nanoparticles even in similar experimental conditions in terms of number of proteins, type of the proteins, protein abundance, etc." | |
| "Our findings suggest that there is a critical need for the development of a standard protocol and workflows for protein corona analysis with regard to LC-MS/MS sample preparation and analysis to minimize the heterogeneity of the final outcomes," he adds. | |
| One of the possible next stages of this research would be developing a standard LC-MS protocol and send it to various core facilities/proteomics centers for characterization of protein corona composition to minimize the heterogeneity of the protein corona outcomes across core facility. | |
| Mahmoudi concedes that developing a unique standard protocol for standard LC-MS analysis and convincing the proteomics core labs to follow the same procedures for protein corona analysis would be a considerable challenge: Different cores facilities and proteomics centers don't have access to the same instrumentation (i.e., mass spectrometers and chromatography systems), same software (such as commercial database search software), nor could/would they comply with following exact protocols (for example using different enzymes for digestion, reducing agents, cleaning protocols, etc.). Moreover, it would also be impossible to measure the protocol compliance level, e.g., incubation times, quality of materials used, etc. | |
| "In the past decade, we have witnessed limited success in clinical translation of therapeutic nanoparticles and one of the main reasons for this limitation is the heterogeneity of the protein corona results in the field due to the complex composition of the protein corona and absence of a standard protocol/approach for characterization and interpretation of the protein corona profiles," Mahmoudi concludes. "Debugging these overlooked factors will result in our deeper understanding of the biological identity of nanoparticles in in vivo environments." | |
 By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
|
|
|
Become a Spotlight guest author! Join our large and growing group of guest contributors. Have you just published a scientific paper or have other exciting developments to share with the nanotechnology community? Here is how to publish on nanowerk.com. |
|
