Photoactivated Localization Microscopy (PALM): Stochastic Single-Molecule Imaging for Nanoscale Biology
What is Photoactivated Localization Microscopy?
Photoactivated Localization Microscopy (PALM) is a super-resolution imaging technique that allows for the visualization of biological structures at the nanoscale. It surpasses the diffraction limit of conventional light microscopy by utilizing the properties of fluorescent proteins or dyes that can be photoactivated or photoswitched. PALM enables the localization and mapping of individual molecules within a sample, providing unprecedented insights into cellular processes and structures.
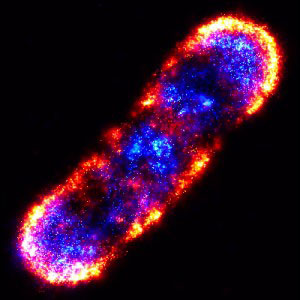
Principles of PALM
PALM relies on the following key principles:
Photoswitchable Fluorescent Labels
PALM employs photoswitchable fluorescent proteins or dyes that can be activated from a dark state to a fluorescent state by exposure to light of a specific wavelength. These labels are genetically fused to the proteins of interest or chemically attached to specific targets within the sample.
Stochastic Activation and Localization
During PALM imaging, a low-intensity activation light is used to stochastically switch on a sparse subset of fluorescent labels within the sample. The activated molecules are then excited with a readout laser and imaged until they photobleach. This process is repeated multiple times, activating and imaging different subsets of molecules in each cycle.
Single-Molecule Localization
The fluorescence emission from individual activated molecules appears as diffraction-limited spots in each image frame. By fitting these spots with a mathematical function, such as a Gaussian function, the precise location of each molecule can be determined with nanometer accuracy. This localization procedure is performed for all the activated molecules in each cycle.
Image Reconstruction
The localized positions of individual molecules from all the imaging cycles are combined to reconstruct a super-resolution image. This image represents the distribution of the labeled molecules within the sample, revealing fine structural details that are not visible in conventional microscopy.
Comparison with Other Super-Resolution Techniques
PALM is one of several super-resolution microscopy techniques that have emerged in recent years. Other notable techniques include:
Stochastic Optical Reconstruction Microscopy (STORM)
stochastic optical reconstruction microscopy is a closely related technique to PALM, utilizing photoswitchable dyes instead of fluorescent proteins. It follows similar principles of stochastic activation, localization, and image reconstruction.
Stimulated Emission Depletion (STED) Microscopy
Stimulated Emission Depletion microscopy achieves super-resolution by using a donut-shaped depletion laser to confine the fluorescence emission to a sub-diffraction volume. It offers high temporal resolution but requires specialized laser systems and fluorophores.
Structured Illumination Microscopy (SIM)
Structured Illumination Microscopy uses patterned illumination to encode high-frequency spatial information into the observed image. By acquiring multiple images with different illumination patterns and processing them, a super-resolution image can be reconstructed. SIM provides a modest resolution improvement compared to PALM and STORM.
Compared to these techniques, PALM offers several advantages:
- High spatial resolution, typically in the range of 10-50 nanometers.
- Compatibility with a wide range of photoswitchable fluorescent proteins and dyes.
- Ability to study live cells and dynamic processes.
- Relatively simple instrumentation compared to techniques like STED.
Applications of PALM
PALM has found numerous applications in biological and biomedical research:
Protein Localization and Clustering
PALM enables the precise mapping of protein distributions within cells, revealing the spatial organization and clustering of proteins at the nanoscale. This information is crucial for understanding protein function, interactions, and signaling pathways.
Membrane Dynamics and Receptor Trafficking
PALM has been used to study the dynamics of membrane proteins, such as receptors and ion channels, with high spatial and temporal resolution. It provides insights into the trafficking, diffusion, and clustering of these proteins, which are essential for cellular communication and signaling.
Chromatin Organization and Gene Expression
PALM has been applied to investigate the nanoscale organization of chromatin in the nucleus, revealing the spatial distribution of histones, transcription factors, and other chromatin-associated proteins. This information is valuable for understanding gene regulation and epigenetic mechanisms.
Challenges and Future Perspectives
Despite its remarkable capabilities, PALM faces several challenges. One limitation is the reliance on photoswitchable fluorescent labels, which may perturb the native structure and function of the target proteins. The development of smaller, more photostable, and less disruptive labels is an active area of research.
Another challenge is the need for specialized image analysis software and computational resources to process the large datasets generated by PALM. Advances in data analysis algorithms and high-performance computing will facilitate the widespread adoption of PALM in biological research.
Future developments in PALM are expected to focus on improving the temporal resolution, enabling the study of fast dynamic processes. The integration of PALM with other imaging modalities, such as electron microscopy and Raman spectroscopy, will provide a more comprehensive understanding of biological systems at multiple scales.
Further Reading
Cold Spring Harbor Protocols, Photoactivated localization microscopy (PALM): an optical technique for achieving ~10-nm resolution
Nature Reviews Methods Primers, Single-molecule localization microscopy
