| Posted: August 20, 2008 |
Spying on self-assembly |
|
(Nanowerk News) Proteins attaching to gold nanoparticles don't mill around randomly, but organise into clusters, according to UK scientists who say they have for the first time spied in detail peptides assembling on a surface.
|
|
The researchers say their study ("Supramolecular Domains in Mixed Peptide Self-Assembled Monolayers on Gold Nanoparticles") is a step towards building complex self-assembled objects at the nanoscale - such as artificial enzymes or other nanomachines that could be used in drug delivery or diagnostics.
|
|
A conventional electron microscope snapshot can't capture in detail the assembly of a small, fast-moving 'soft' biological material on a surface - such as a protein shuffling around on a nanoparticle.
|
|
But a team led by Raphaël Lévy at the University of Liverpool have now glimpsed the self-assembly process with a technique used to study protein complexes. The researchers covered gold nanoparticles with a monolayer of self assembling peptides. Most of these were short, but a few were grafted on to longer chains to create structures around 20 amino acids long - in effect creating functionalised peptide 'trees' amongst a layer of 'grass' shuffling over a gold nanoparticle.
|
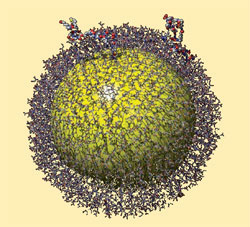 |
| Glimpsing how peptides self-assemble on a gold nanoparticle
|
|
Lévy's team then measured the distance between peptide trees on the surface by adding a cross-linking probe molecule, both ends of which could bind to pre-arranged sites on the long peptides.
|
|
The 1 nm cross-linker joined peptide strands together: if the peptides were isolated they looped around on themselves, whereas if they were close together, bridges formed between them. By testing different concentrations of peptides on nanoparticles - and using gel electrophoresis to measure the ratios of loops to bridges when a probe was added - the team measured the average distance between peptides.
|
|
The researchers were surprised to find that much more self-organisation was present than they had expected. Even with only a few peptide chains on a nanoparticle 10nm in diameter, the team observed far more bridges than their calculations predicted, in some cases spotting dimers and trimers forming.
|
|
'As soon as you have three or more peptides on the surface, you start to see bridges between them,' Lévy says. 'It seems that once a long peptide has anchored on the surface, others are more likely to anchor or move close to it.'
|
|
'From a practical point of view we could use this technique to engineer specific nanostructures on the surface in future,' says David Fernig, a co-author on the project. 'It can also help us to better understand processes where molecules self-organise into structures - such as protein folding.'
|
|
'Nanoparticle surfaces are dynamic environments where the unique balance of entropy and enthalpy can provide surface architectures ranging from highly ordered to completely random,' says Victor Rotello, who works on similar systems at the University of Massachusetts in Amherst, US. 'This demonstration of preorganisation is of great importance in the design of nanoparticle based delivery vehicles, sensors, and devices.'
|

