| Posted: Sep 04, 2007 | |
Sucking nanospaghetti through nanopores - the art of single-molecule spectroscopy |
|
| (Nanowerk Spotlight) There is a significant and growing need across the research and medical communities for low-cost, high throughput DNA separation and quantification techniques. The isolation of DNA is a prerequisite step for many molecular biology techniques and experiments. Although single molecule techniques afford extremely high sensitivity, to date, such experiments have remained within the confines of academic and research laboratories. The primary reasons for this state of affairs relate to throughput, detection efficiencies and analysis times. For example, in a conventional solution-based single molecule detection experiment, one can only detect approximately 10,000 molecules per minute, or one molecule every 6 milliseconds. While this may sound a lot, consider that a small drop of water (ca. 5 ml) contains approx. 1.67 x 1023 molecules, that is 1.67 followed by 23 zeros. At that speed you need over 100 trillion years to detect all the water molecules in this single drop. Using a novel nanopore array developed by researchers in the UK, expect to be able to detect up to 1 million molecules simultaneously in the same 6 millisecond time window (and bringing the timeframe for analyzing the molecules in a single water drop down to some 60 billion years – about five to six times the estimated age of the universe). | |
| The above example exaggerates a bit, of course. Compared to a water molecule, which is very small (it consists of only three atoms with an overall diameter of less than 0.3 nm), a DNA molecule is very large. In the real world you would never analyze a drop of DNA but much, much smaller quantities, but even then existing methods are considered to be unacceptably slow. Although it has been seven years that the completion of the rough draft of the human genome was announced, much effort is still focused on identifying genes responsible for specific biological functions (or diseases) and determining the DNA sequence bearing the information. | |
| "In recent years, the creation of nanochannels or nanopores in thin membranes has attracted much interest due to the potential to isolate and sense single DNA molecules while they translocate through the highly confined channels" Dr. Joshua Edel, a lecturer in micro- and nanotechnology at the Imperial College London, explains to Nanowerk. "Nanopores for such applications have already been fabricated but in all studies to date, the detection of translocation events is performed electrically by measuring the ionic current" (what this means is that molecules translocating through a nanopore will momentarily perturb the ionic current, with the duration of the perturbation and the magnitude of the current blockade providing more detailed information about molecular shape and structure). | |
| Edel's group, together with collaborators from Drexel University, recently presented proof-of-concept studies that describe a novel approach for optically detecting DNA translocation events through an array of solid-state nanopores which allows for ultrahigh-throughput, parallel detection at the single-molecule level ("Single-Molecule Spectroscopy Using Nanoporous Membranes"). | |
 |
|
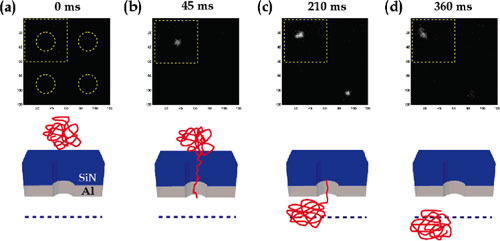
| Fluorescence image of two DNA translocation events occurring under an applied voltage of 0.45 V at t=0 (a), t=45 ms (b), t=210 ms (c), and t=360 ms (d). Each pixel represents an area of 81 x 81 nm2. The dotted circles in frame (a) indicate the location of the pores. Illustrations below each image frame provide an indication of the progression of DNA through the pore and the illumination plane (dotted line). (reprinted with permission from American Chemical Society) | |
| The above figure shows a sequence of images of typical translocation events for four pores as captured by a CCD camera. Figure (b) illustrates the onset of a translocation event in the top left hand pore. DNA passage through the field of view is complete after approximately 360 ms. | |
| "The single molecule studies performed by Guillaume Chansin, a PhD student, within my research group are very exciting and novel from a technological perspective" says Edel. "Firstly, this work is the first demonstration of using fluorescence detection to monitor translocation events within a nanopore array. Secondly, this is the first true demonstration of an approach leading towards high throughput single molecule detection confined within nanofluidic structures." | |
| This technology functions by electrokinetically driving DNA strands through sub-micrometer-sized holes (in this case 300 nm) on an aluminum/silicon nitride membrane. During the translocation process, the molecules are confined to the walls of the nanofluidic channels, allowing 100% detection efficiency. Importantly, the opaque aluminum layer acts as an optical barrier between the illuminated region and the analyte reservoir. In these conditions, high-contrast imaging of single-molecule events can be performed. | |
| Edel notes that the majority of work to date using nanopores utilizes ionic blockage currents to monitor translocation events. "Unfortunately, simply measuring blockage currents can only be performed in a single pore" he says. "One of our motivations to using optical detection was to ensure we can probe multiple holes simultaneously allowing for true high throughput detection." | |
| "Our results indicate that it is possible to obtain high spatial resolution DNA analysis while independently controlling the applied voltage that drives the molecules into the nanopore" says Edel. "A critical feature of the generic approach is the possibility of parallelizing molecular analysis by probing an entire array of nanopores under uniform illumination. | |
| "There are plenty of potential applications ranging from DNA sequencing, fragment sizing, sieving, separations, and rare event diagnostics" says Edel. "For example, using analytical technologies that exist today it is essentially impossible to detect a single DNA strand within a standard blood sample (of a few milliliters) within a reasonable time frame. The technology we describe can potentially allow for such detection to be performed both rapidly and efficiently." | |
| It seems that nanopore research is an exciting and growing field. Expect optical detection to play a dominant role in the future of this technology. | |
 By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
|
Become a Spotlight guest author! Join our large and growing group of guest contributors. Have you just published a scientific paper or have other exciting developments to share with the nanotechnology community? Here is how to publish on nanowerk.com.
