Showing Spotlights 929 - 936 of 2785 in category All (newest first):
 Inspired by the multiple water purification mechanisms of water hyacinth, a group of researchers has combined different decontamination techniques - adsorption, photocatalytic degradation, distillation - into a single paper-based composite system for water purification by using solar light as the clean energy input. Compared with water treatment with a single mechanism, the multifunctional composite reported in this work could significantly enhance the clean water generation efficiency and maximize the use of the broad-spectrum solar light in a facile and effective way.
Inspired by the multiple water purification mechanisms of water hyacinth, a group of researchers has combined different decontamination techniques - adsorption, photocatalytic degradation, distillation - into a single paper-based composite system for water purification by using solar light as the clean energy input. Compared with water treatment with a single mechanism, the multifunctional composite reported in this work could significantly enhance the clean water generation efficiency and maximize the use of the broad-spectrum solar light in a facile and effective way.
Jun 14th, 2016
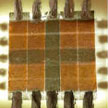 Using multilayer graphene as an electrically reconfigurable optical medium, researchers have demonstrated an optoelectronic framework compatible with conventional printing paper. The device consist of two multilayer graphene layers transfer-printed on both sides of the paper. In this configuration, multilayer graphene simultaneously operates as the electrically reconfigurable optical medium and electrically conductive electrodes. In addition, the paper substrate yields a flexible and foldable mechanical support for the graphene layers and it holds the electrolyte in the network of hydrophilic cellulose fibers.
Using multilayer graphene as an electrically reconfigurable optical medium, researchers have demonstrated an optoelectronic framework compatible with conventional printing paper. The device consist of two multilayer graphene layers transfer-printed on both sides of the paper. In this configuration, multilayer graphene simultaneously operates as the electrically reconfigurable optical medium and electrically conductive electrodes. In addition, the paper substrate yields a flexible and foldable mechanical support for the graphene layers and it holds the electrolyte in the network of hydrophilic cellulose fibers.
Jun 8th, 2016
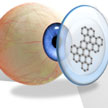 The latest example of a graphene-based wireless sensor that could make 24-hour healthcare easier to achieve by enabling wireless monitoring of various biomedical events in order to gain a more comprehensive assessment of the wearer's healthcare status. This novel device, which detects chemical/molecular agents and lengths of exposure, can be used as lightweight and transparent wearable or bio-implantable electronic sensor. It may provide an inexpensive way to detect in real-time the biomedical of interest.
The latest example of a graphene-based wireless sensor that could make 24-hour healthcare easier to achieve by enabling wireless monitoring of various biomedical events in order to gain a more comprehensive assessment of the wearer's healthcare status. This novel device, which detects chemical/molecular agents and lengths of exposure, can be used as lightweight and transparent wearable or bio-implantable electronic sensor. It may provide an inexpensive way to detect in real-time the biomedical of interest.
Jun 6th, 2016
 In view of the scientific and technological potential of CNTs, it is of immense importance to know who should be credited for their discovery. In the present article, we have made an attempt to give a glimpse into the discovery and early history of this fascinating material for our readers. Carbon nanotubes possess unique combination of extraordinary mechanical, electronic, transport, electrical and optical, properties and nanoscale sizes making them suitable for a variety of applications ranging from engineering, electronics, optoelectronics, photonics, space, defence industry, medicine, molecular and biological systems.
In view of the scientific and technological potential of CNTs, it is of immense importance to know who should be credited for their discovery. In the present article, we have made an attempt to give a glimpse into the discovery and early history of this fascinating material for our readers. Carbon nanotubes possess unique combination of extraordinary mechanical, electronic, transport, electrical and optical, properties and nanoscale sizes making them suitable for a variety of applications ranging from engineering, electronics, optoelectronics, photonics, space, defence industry, medicine, molecular and biological systems.
Jun 3rd, 2016
 Over the past few decades, the development of electron microscopy has gone hand in hand with techniques for atomically precise fabrication of 3D structures based on electron and ion beams. A recent review article illustrates the use of focused electron and ion beams (e-beams and i-beams) to induce highly localized chemical reactions at solid-vapor and solid-liquid interfaces, amorphous to crystalline phase transformations with atomic layer precision, and the motion of specific single dopant atoms within crystal lattices, thus laying the foundation for atomically precise directed assembly of materials and devices.
Over the past few decades, the development of electron microscopy has gone hand in hand with techniques for atomically precise fabrication of 3D structures based on electron and ion beams. A recent review article illustrates the use of focused electron and ion beams (e-beams and i-beams) to induce highly localized chemical reactions at solid-vapor and solid-liquid interfaces, amorphous to crystalline phase transformations with atomic layer precision, and the motion of specific single dopant atoms within crystal lattices, thus laying the foundation for atomically precise directed assembly of materials and devices.
May 31st, 2016
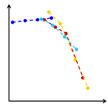 In the fields of toxicology and ecotoxicology, doses are commonly expressed in weight concentration for non-soluble compounds because this is very convenient experimentally. However, when it comes to nanoparticles, the weight of the nanomaterial is not a relevant parameter, especially when it is required to compare different kinds of nanoparticles because their density is very different. Researchers have now shown that the usual approach based on mass concentrations fails to compare the toxicities of different engineered carbon nanoparticles.
In the fields of toxicology and ecotoxicology, doses are commonly expressed in weight concentration for non-soluble compounds because this is very convenient experimentally. However, when it comes to nanoparticles, the weight of the nanomaterial is not a relevant parameter, especially when it is required to compare different kinds of nanoparticles because their density is very different. Researchers have now shown that the usual approach based on mass concentrations fails to compare the toxicities of different engineered carbon nanoparticles.
May 24th, 2016
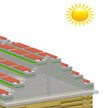 Self-powered nanotechnology based on these nanogenerators aims at powering nanodevices and nanosystems using the energy harvested from the environment in which these systems are suppose to operate. An interesting approach comes from a group of Chinese scientists who propose to scavenge the large amounts of wasted wind energy in cities: they propose hybridized nanogenerator that consists of a solar cell and a triboelectric nanogenerator, which can be utilized to individually or simultaneously scavenge solar and wind energies.
Self-powered nanotechnology based on these nanogenerators aims at powering nanodevices and nanosystems using the energy harvested from the environment in which these systems are suppose to operate. An interesting approach comes from a group of Chinese scientists who propose to scavenge the large amounts of wasted wind energy in cities: they propose hybridized nanogenerator that consists of a solar cell and a triboelectric nanogenerator, which can be utilized to individually or simultaneously scavenge solar and wind energies.
May 23rd, 2016
 High-temperature heaters, such as furnaces, are widely used in chemical reactions, materials synthesis and device processing. The limitations of these heating devices often are their bulky size, weight, low maximum heating temperatures and slow ramp rates. To overcome these limitations, and to provide a heating element with a high temperature range to the target object in a micro- and nanoscale environment, researchers have developed a 3D-printable high-temperature, high-rate heater that can be applied to a wide range of nanomanufacturing when precise temperature control in time, placement, and the ramping rate is important.
High-temperature heaters, such as furnaces, are widely used in chemical reactions, materials synthesis and device processing. The limitations of these heating devices often are their bulky size, weight, low maximum heating temperatures and slow ramp rates. To overcome these limitations, and to provide a heating element with a high temperature range to the target object in a micro- and nanoscale environment, researchers have developed a 3D-printable high-temperature, high-rate heater that can be applied to a wide range of nanomanufacturing when precise temperature control in time, placement, and the ramping rate is important.
May 19th, 2016
 Inspired by the multiple water purification mechanisms of water hyacinth, a group of researchers has combined different decontamination techniques - adsorption, photocatalytic degradation, distillation - into a single paper-based composite system for water purification by using solar light as the clean energy input. Compared with water treatment with a single mechanism, the multifunctional composite reported in this work could significantly enhance the clean water generation efficiency and maximize the use of the broad-spectrum solar light in a facile and effective way.
Inspired by the multiple water purification mechanisms of water hyacinth, a group of researchers has combined different decontamination techniques - adsorption, photocatalytic degradation, distillation - into a single paper-based composite system for water purification by using solar light as the clean energy input. Compared with water treatment with a single mechanism, the multifunctional composite reported in this work could significantly enhance the clean water generation efficiency and maximize the use of the broad-spectrum solar light in a facile and effective way.
 Subscribe to our Nanotechnology Spotlight feed
Subscribe to our Nanotechnology Spotlight feed





