| Posted: Nov 01, 2017 | |
Virtual cell model predicts cell response to substrate topography (w/video) |
|
| (Nanowerk Spotlight) The crucial roles of the physicochemical properties of cell culture substrates on function and behavior of a wide range of the cells are becoming well-studied in the current literature, using experimental approaches. However, development of in silico approaches for prediction of cell responses to the physicochemical properties of substrates is still in its infancy. | |
| In new work, reported in ACS Nano ("Development of a Virtual Cell Model to Predict Cell Response to Substrate Topography"), an international team of researchers has developed a unifying computational framework to create a multi-component virtual cell model to probe cell function/behavior in silico. | |
| By correlation their modeling data with experimental outcomes, they demonstrated the capability of this model to predict cell behavior outcomes, i.e. to predict changes in whole cell and cell nucleus characteristics – in terms of shape, direction, and even chromatin conformation – on a range of cell substrates. | |
| "Using the virtual cell approach, we can easily track the variation of cell and nucleus shapes together with chromatin conformational changes during differentiation," Morteza Mahmoudi, Director of and Principal Investigator at the NanoBio Interactions Laboratory at Tehran University of Medical Sciences, tells Nanowerk. "Our findings should help researchers understand the mechanisms involved in shape-induced physical differentiation of stem cells." | |
| He points out that the virtual cell approach may provide a reliable, efficient, and fast high-throughput approach for the development of optimized substrates for a broad range of cellular applications including stem cell differentiation. | |
 |
|
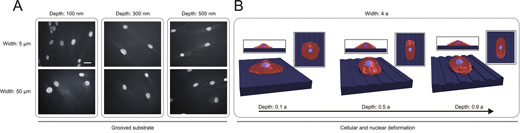
| Prediction of deformation and elongation of the cells on the grooved substrate using a “virtual cell modeling”. (A) Cultivated cells on grooved substrates with widths of 5 and 50 µm and depths of 100-400 nm. Scale bars indicate 25 µm. (B) Morphological alteration of the virtual cell on the grooved substrate with a width of 4a (i.e., a is the reduced length unit) and a depth of 0.1a - 0.9a. (Reprinted with permission by American Chemical Society) (click on image to enlarge) | |
| The prediction of cell behavior, using computational approaches, could be very powerful in speeding up the design of optimal stem cell culture substrates. | |
| Mahmoudi adds that the proposed model has the capacity not only to predict shape and conformation of the cells qualitatively but also to give quantitative results if the adequate and proper parameters are used. | |
| "Consequently, the findings should help researchers understand the mechanisms involved in shape-induced physical differentiation of stem cells," he notes. | |
| Previously, the team has used their cell-imprinted substrates to demonstrate reliable and efficient control of MSC differentiation toward | |
|
|
|
| They have also demonstrated that cell-patterned substrates modulate the differentiation, redifferentiation, and transdifferentiation of a variety of cells. | |
| In this new study, the scientists' goal was to probe whether the virtual cell can show the mechanism behind the observed experimental results with regard to the physical differentiation. | |
| "Our main hypothesis in the experimental approach was that the stem cells cultured on the cell-imprinted substrates are driven to adopt the specific shape and molecular characteristics of the mature cell types that had been used as a template for the cell-imprinting process," says Mahmoudi. "Our in silico results verified this hypothesis, as the virtual cell revealed that the cultured cells can mimic the shape of the imprinted substrates and change their geometry in both cellular and nuclear structures." | |
| "In addition, we use our models to predict that the confining geometry of the imprints has effects on the chain arrangement of simulated chromatin fibers in the nuclei, which may have roles in the physical differentiation process." | |
| The movie shows top, side and 3D views of a virtual cell on grooved substrate becoming elongated along a grooved substrate with elongation increasing with direct proportionality to increased groove depth. | |
| It is important to note that the authors' goal in this paper is to provide qualitative behavior of the MSCs. By using the real parameters (e.g., Young’s modulus and the force generated by actin polymerization) of a particular cell type in experiments, this model could be potentially customized for other specific cell types to provide quantitative outcomes. | |
| "The optimal substrates, for directing specific cell function (e.g., differentiation), will have translational impact through the provision of appropriate substrates to target desired stem cell phenotypes," Mahmoudi notes. "We propose that they will find roles in prevention/prediction of diseases through better in vitro drug screening and through helping the cell supply chain to underpin effective cell therapy." | |
| "Our results provide evidence that the virtual cell model could be employed as a platform to understand how cells sense and respond to the extracellular matrix," heconcludes. "But, it is noteworthy that the process of cell differentiation is a complex and dynamic phenomenon; to further enhance the predictability of our virtual cell model, the model should be further improved to consider the incubation time and its effects on the differentiation process." | |
 By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
|
|
|
Become a Spotlight guest author! Join our large and growing group of guest contributors. Have you just published a scientific paper or have other exciting developments to share with the nanotechnology community? Here is how to publish on nanowerk.com. |
|
