| Jun 17, 2019 | |
Early cancer detection with sensory protein corona 'fingerprints' |
|
| (Nanowerk Spotlight) When nanoparticles enter the blood stream or other biological fluids, they are quickly surrounded by a layer of proteins. This so-called protein corona on the nanoparticles hinders interactions between the targeting ligands on the nanoparticles and their binding partners on the cells' surface. | |
| Previous research findings in 2014, reported by Mahmoudi's Lab, have indicated that different individuals may have a personalized protein corona owing to their distinct type, severity and period of disease, heterogeneity, individual genetic variations and environmental factors. This means that the type of disease has a crucial role in the protein composition of the nanoparticle corona and that exactly the same nanoparticles may have different therapeutic and/or toxic impacts on different individuals. | |
| By combining the concepts of disease-specific protein corona and sensor array technology, researchers now have created a label-free platform for the early detection and identification of diseases. | |
| They report their findings in Nanoscale Horizons ("Disease-specific protein corona sensor arrays may have disease detection capacity"). | |
| "The possibility to measure panels of specific and selective biomarker proteins has the potential to revolutionize cancer screening, detection and monitoring," Morteza Mahmoudi, an Assistant Professor at Brigham and Women’s Hospital, Harvard Medical School, tells Nanowerk. "Our protein corona sensor array, rather than detecting a specific biomarker, provides pattern recognition of the corona protein composition adsorbed on the liposomes. By selecting disease-specific corona profiles of different nanoparticles, using supervised classifiers, we created a unique protein pattern, which was the 'fingerprint' of each type of cancer." | |
 |
|
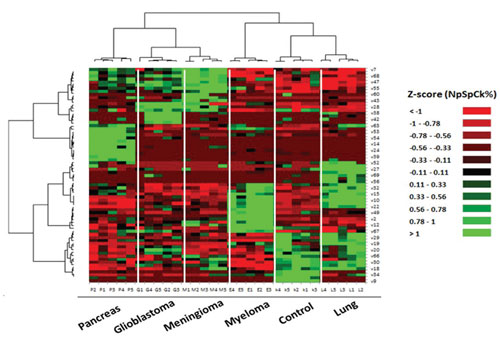
| Identification and discrimination of cancers using protein corona sensor arrays. 51 proteins identified as capable of distinguishing among the six groups are presented in a ‘Heat Map’ generated using an unsupervised cluster algorithm. Visual inspection of the heat map, based on the raw data of 69 important markers, demonstrates cancer-specific protein corona signature and clear clustering of six groups of samples (five groups of cancerous samples plus normal samples) and also expected similarities among five patients from each group. (Reprinted with permission by Royal Society of Chemistry) (click on image to enlarge) | |
| The results obtained by Mahmoudi and an international collaboration of researchers revealed that although no single protein corona composition from a single nanoparticle provides this 'fingerprint' feature, the pattern of corona composition derived from the nanoparticle sensor array provides a unique 'fingerprint' for each type of cancer. | |
| As a feasibility study, the team focused on five distinct human cancers (lung cancer, glioblastoma, meningioma, myeloma, and pancreatic cancer) as a model disease ex vivo. | |
| The sensor array is composed of three different cross-reactive liposomes with various lipid compositions: anionic liposomes, cationic liposomes, and zwitterionic liposomes. The researchers characterized the protein corona profiles at the surface of these liposomes by nano liquid chromatography tandem mass spectrometry after exposure to the plasma of patients diagnosed with each of five cancers. It is obvious that by increasing the sensor array elements (i.e., nanoparticles' type and their physico-chemical characteristics), the disease detection accuracy of the protein corona sensor array technology will be enhanced. | |
| "To probe the capacity of this platform for very early detection of cancers, we used cohort plasma obtained from healthy people who were later diagnosed with lung, pancreas, and brain cancers several years after plasma collection and the outcomes revealed that the approach could identify and discriminate the cancers," Mahmoudi explains. | |
| "Our results suggest that the disease-specific protein corona sensor array will not only be instrumental in the screening, detection, and identification of diseases, but may also help identify novel protein pattern markers whose role in disease development and/or disease biology has not been appreciated so far," he concludes. | |
 By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
|
|
|
Become a Spotlight guest author! Join our large and growing group of guest contributors. Have you just published a scientific paper or have other exciting developments to share with the nanotechnology community? Here is how to publish on nanowerk.com. |
|
