| Posted: Apr 02, 2007 | |
Native protein nanolithography that can write, read and erase |
|
| (Nanowerk Spotlight) Proteins are very specific about which other proteins or biochemicals they will interact with and therefore are of great use for biosensing applications. For instance, if a malignant cancer develops in the human body, the cancer cells produce certain types of proteins. Identifying such proteins enables early detection of cancer. | |
| One of the goals of nanobiotechnology is to develop protein chips that are sensitively responsive to a very tiny amount of specific proteins in order to enable such early stage diagnosis. For example, a protein that is known to bind to a protein produced by a cancer cell could be attached to a biochip. If this particular cancer cell protein were present in a sample passed over the chip, it would bind to the protein on the chip, causing a detectable change in the electrical signal passing through the chip. This change in the electrical signal would be registered by the device, confirming the presence of the protein in the sample. | |
| While this sounds very promising for the future of diagnostic systems, with the promise of protein chips capable of single-molecule resolution, the controlled assembly of proteins into bioactive nanostructures still is a key challenge in nanobiotechnology. | |
| Researchers in Germany took a further step towards this goal by developing a native protein nanolithography technique that allows for the nanostructured assembly of even fragile proteins. | |
| Existing nanolithographic techniques are operated under vacuum or ambient atmosphere conditions – settings which are not compatible with most biological molecules, as oxidation, dehydration, and organic solvents impair these delicate entities. Being able to perform direct patterning with proteins therefore has been the exception and maintaining their bioactivity is only possible in case of rather stable proteins such as antibodies. For this reason, the biochemically interesting proteins such as receptors from signal transduction pathways or large macromolecular complexes have not yet been included in protein chip fabrication. | |
| "We recognized a technological gap in this field of protein nanopatterning and aimed to develop a process that protects biomolecules from non-physiological conditions" Ali Tinazli explains to Nanowerk. "The technique we came up with, native protein nanolithography (NPNL), has a very straightforward process control and it has been transferred easily between different instruments and labs. Standard atomic force microscopy (AFM) equipment is fully sufficient for this purpose." | |
| Tinazli is the first author of a recent paper in Nature Nanotechnology ("Native protein nanolithography that can write, read and erase") that describes a nanolithography technique that permits rapid writing, reading and erasing of protein arrays in a versatile manner. Developed by a group of researchers from the Cellular Biochemistry lab of Professor Robert Tampé at the Johann-Wolfgang-Goethe University of Frankfurt, and the Max-Planck-Institute of Biochemistry near Munich, the corresponding protein chip platform is suitable for any His-tagged proteins (an amino acid motif in proteins used to detect protein-protein interactions). | |
 |
|
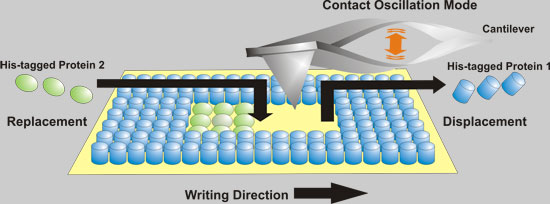
| Fabrication of rewritable protein nanoarrays on SAMs by native protein nanolithography. (Image: Ali Tinazli) | |
| The researchers in Germany based their novel and easy-to use fabrication technique for protein nanoarrays on metal-chelating self-assembled monolayers combined with AFM-based nanolithography. Affinity-captured proteins are mechanically displaced in a special patterning mode of AFM. For lateral structuring of multiple protein assemblies, these proteins are replaced simultaneously or sequentially by different His-tagged proteins. | |
| What is so special about this AFM technique is that it permits instant switching between imaging and replacement of immobilized proteins in their native state without any change of the setup or the tip. In NPNL the first self-assembled protein layer acts as a biocompatible and ductile patterning material (protein resist). | |
| Affinity-captured proteins can be replaced by applying the AFM tip in contact oscillation mode (a term the researchers coined for a vibrational mode where the AFM tip is in constant contact with the sample), and the generated structures can be erased and refilled specifically, with different proteins, in a uniform and functional manner. | |
| "The novelty of our approach is that it allows the fabrication of bioactive protein nanoarrays down to a resolution of 50 nm in a fast and versatile manner without the need of vacuum or ambient atmosphere conditions – but under conditions of a physiological solution" says Tinazli. NPNL serves requirements in nanobiotechnology, where physiological ambient conditions, such as in aqueous solution, are highly desirable for ensuring the preservation of biological functionality during and after array fabrication." | |
| This new technique enables the fabrication of rewritable protein nanoarrays, allowing a previously unobtainable flexibility in nanolithography and experimentation in biosensing and single-molecule studies. | |
| The researchers also demonstrated that this erase-and-write (i.e. 'displacement' and 'replacement') technique allows a more complex lateral organization of protein assemblies in multiplexed arrays, consisting of a series of different proteins and protein complexes in a unique orientation. | |
| Tinazli points out that this higher degree of multiplexity of protein nanoarrays will be key in the next steps the researchers are planning to take. "We will spend our future effort on the fabrication of nanocatalytic centers and highly multiplexed protein assembly lines" he says. | |
| Specific applications of this work will facilitate the design of protein arrays for biosensors, single-molecule studies, and medical diagnostics. Especially applications in the biosensor field, in combination with nanooptical biosensing elements, will provide new research tools allowing insights into intermolecular interactions. | |
 By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
|
Become a Spotlight guest author! Join our large and growing group of guest contributors. Have you just published a scientific paper or have other exciting developments to share with the nanotechnology community? Here is how to publish on nanowerk.com.
