| Posted: Nov 21, 2016 | |
From gene sequencing to genome mapping with nanopores and nanochannels |
|
| (Nanowerk Spotlight) One reason why people are so excited about nanopore DNA sequencing is that the technology could possibly be used to create 'tricorder'-like devices for detecting pathogens or diagnosing genetic disorders rapidly and on-the-spot. | |
| Nanopores function as membrane channels in all living systems, where they serve as sensitive electromechanical devices that regulate electrical potential, ionic flow, and molecular transport through the cell membrane. Scientists are studying nanopore construction with the goal of making man-made cell membranes and single molecule detectors. Biological protein pores used in sequencing have a diameter of about 1-1.5 nm. | |
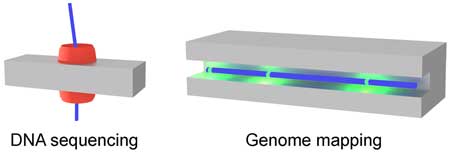
| Sequencing technologies have made it cheaper and faster to read the sequence of bases on a strand of DNA. A promising technology to take these advances further is nanopore sequencing. Individual strands of DNA are moved through a nanopore gap not much wider than the DNA itself. As the DNA passes through the nanopore, continuous information is gained about the sequence of individual bases – the A, C, G and Ts that make up DNA (read more in our previous Nanowerk Spotlight: "DNA sequencing reaches new lengths"). | |
| Yet, nanopores enable another important way to analyze DNA: genomes can also be mapped. In particular, chromosomal DNA strands are placed inside narrow nanochannels of microscale length to check for genetic rearrangements such as insertions and deletions. Genome mapping aims to identify the location of genes on chromosomes and their relative distance. | |
 |
|
| In a Perspective article by Stefan Howorka at University College London and Zuzanna S. Siwy at University of California, Irvine, also published in ACS Nano ("Nanopores and Nanochannels: From Gene Sequencing to Genome Mapping"), the authors describe the principle, advantages, and challenges of genome mapping with nanochannels and relate it to DNA sequencing. | |
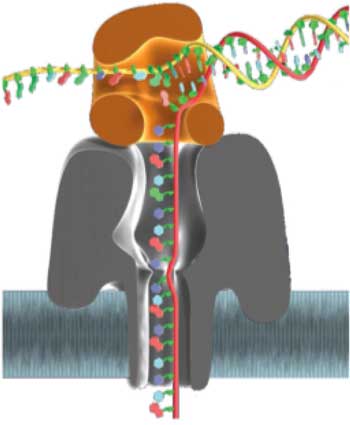
| To obtain sequence information with nanopores, individual strands are moved along an internal reading head. The reading head is a narrow and thin molecular constriction of ca. 1.2 nm that is composed of several amino acid residues. When the DNA strand passes by this constriction, bases can be identified by monitoring the ionic current flowing through the channel. | |
 |
|
| DNA sequencing with nanopores. A protein pore is equipped with an enzyme motor, which threads the DNA strand base-by-base. The readout is based on transmembrane current that is modulated by the presence of one or only a few bases in the narrowest part of the pore. (© Nature) | |
| One important feature of nanopore sequencing is that it can cover read lengths of up to 200,000 base pairs (bp), compared to a range of up to 1000 bp for existing ensemble sequencing methods. However, as the authors point out, a challenge is that strings of identical bases with different repeat numbers cannot be well discriminated. | |
| A novel genome mapping approach highlighted in a recent paper (ACS Nano, "Super-Resolution Genome Mapping in Silicon Nanochannels") analyzes with nanochannels long chromosomal DNA, in the range of a million base pair length. | |
 |
|
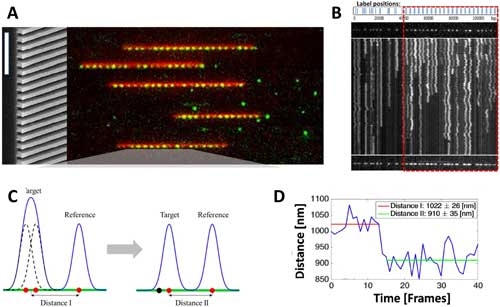
| Genome mapping with nanochannels. (A) Genomic mapping utilizes nanochannels to singulate and to extend single DNA molecules labeled with fluorescent dyes. The molecules are analyzed by microscopy to localize fluorescent spots informative on genomic repeats and deletions. (B) Accuracy of spot localization in genomic mapping is increased by recording a long series of subsequent DNA positions that result from thermal fluctuations. Time advances on the y-axis from top to bottom; the subsequent frames were recorded with 40 ms intervals. (C,D) Two fluorescent dyes separated by a distance below the diffraction limit can be resolved using a combination of stable and blinking fluorophores. Blinking of the dyes resulted in significant changes in the recorded distance from the stable fluorophore. (© American Chemical Society) (click on image to enlarge) | |
| This work makes an important step forward as it tackles the issue of insufficient resolution via a strategy that can achieve a 15-fold improvement. The strategy starts by addressing the main culprit of low resolution, namely, thermal fluctuations of DNA molecules, which is, in turn, a consequence of analyzing DNA strands in solution as opposed to surface-bound DNA. | |
| To mitigate thermal fluctuations, the scientists first employed single-fluorophore imaging of labeled DNA. Single-fluorophore imaging is widely used in the life sciences, but in this work it is successfully applied for genomic mapping in nanochannels. By combining the highly photostable fluorophores with less stable, blinking dyes, the multiple fluorophores could be resolved even though the distance between the dyes was smaller than the diffraction limit. | |
| "It would be advantageous to simplify the design of the current analysis system to make it fully portable," conclude Siwy and Howorka. "Hand-held sequencing devices enable researchers to perform DNA analyses at remote locations and independent of a centralized facility. Portable DNA sequencing and mapping have great potential to facilitate personalized medicine as well as agriculture and homeland security." | |
| Wider adoption of nanopore and nanochannel-based devices would also blur the historic distinction between sequencing and mapping; the very long sequencing reads from nanopores help map larger genomic regions, whereas higher-resolution genomic mapping could provide the sufficiently detailed sequence information that is required for some diagnostic applications. These exciting prospects will undoubtedly fuel more research into the fundamental science and applications of nanopore- and nanochannel-based DNA analysis. | |
 By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
|
|
|
Become a Spotlight guest author! Join our large and growing group of guest contributors. Have you just published a scientific paper or have other exciting developments to share with the nanotechnology community? Here is how to publish on nanowerk.com. |
|
