| Posted: Aug 12, 2009 | |
Towards electronic-based single-molecule DNA sequencing |
|
| (Nanowerk Spotlight) It its more than 25 years of existence, Scanning Tunneling Microscopy has predominantly brought us extremely detailed images of matter at the molecular and atomic level. The Scanning Tunneling Microscope (STM) is a non-optical microscope that scans an electrical probe over a surface to be imaged to detect a weak electric current flowing between the tip and the surface. It allows scientists to visualize regions of high electron density and hence infer the position of individual atoms and molecules on the surface of a lattice. | |
| In nanoelectronics, using single molecules as electronic components is the ultimate goal for future electronic nanotechnology devices and scientists have already made important progress by contacting a single molecule using a STM in a liquid environment at room temperature (read: The long road towards single molecule nanotechnology electronics). | |
| In chemistry, STMs have already been used for real-time single-molecule imaging of an entire chemical reaction. Now, researchers in Japan have managed to partially sequence a single DNA molecule with a STM. Reporting their findings in Nature Nanotechnology ("Partial sequencing of a single DNA molecule with a scanning tunnelling microscope") they show that it is possible to sequence individual guanine bases in real long-chain DNA molecules with high-resolution scanning tunneling microscope imaging and spectroscopy. | |
| Tomoji Kawai, a professor at the Institute of Scientific and Industrial Research at Osaka University, and research assistant Hiroyuki Tanaka have developed a method for extending and fixing DNA strands – a significant step towards the realization of electronic-based single-molecule DNA sequencing. | |
| "Of the four bases, we were able to precisely identify guanine because the STM is able to pick up on the characteristics of its electronic state, which is largely independent of the adsorption structure," says Kawai. "If vibrational spectroscopy is performed using inelastic electron tunneling spectroscopy, it should be possible to identify all of the base molecules. Furthermore, because STM can select a specific position of interest along a DNA strand, the technique could have a unique advantage in analyzing, for example, single nucleotide polymorphisms." | |
| In the past, when scientists tried to sequence single long-chain DNA molecules with the STM some of the difficulties they encountered were due to the unsuitability of the sample preparation methods. Very few studies have been able to make use of the full resolving power of the STM to achieve clear and reproducible observation of individual nucleotides. | |
| Whereas detailed STM imaging, spectroscopy and manipulation studies of molecules are almost always restricted to molecules that can be deposited onto a surface from the gas phase, Kawai and Tanaka have developed a method (for which they have applied for and received a Japanese patent: Kokoku 2005-46665) for stretching out and fixing single-stranded M13mp18 DNA molecules (the double-stranded, covalently closed, circular form of DNA derived from bacteriophage M13, which contains 7,249 bases) by using the flow effects resulting from the oblique injection of a DNA solution onto a substrate using a pulse-injection technique. | |
| "Using this extended DNA, and to check whether or not it is possible to assign the individual guanine units, we measured topography images and dI/dV map images over a 100-nm-wide region" says Kawai. "In the resulting topography image, the individual nucleotides are shown as bright points, which are exceptionally bright in some places." | |
|
|
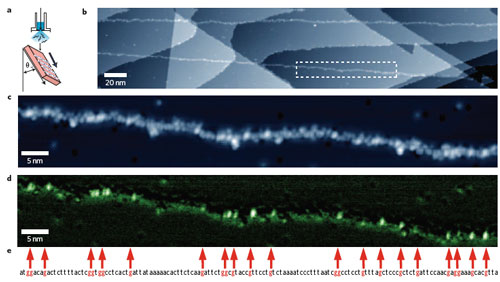
| Deposition and STM analysis of single-stranded M13mp18 DNA molecules on a Cu(111) surface. a) Schematic illustration of the oblique injection method. DNA strands tend to align more perpendicular than horizontal to the flow (injection) direction. The Cu(111) substrate is inclined at ∼45° to the aqueous solution of DNA, which is introduced from a pulse valve. b) Typical wide-area image of M13mp18. Atomic steps in the Cu(111) substrate form a staircase surface structure. In this image, sections of M13mp18 are visualized as linear adsorbed material running from the top left to bottom right. c) An enlarged view of the rectangular region enclosed by the white dashed line in b. d) A dI/dV map of the same region as in c. To maximize the detection of the density of states of guanine, the measurements were made under slightly lower bias conditions than in c. e) Part of the base sequence of M13mp18 obtained from a databank (the sequence of bases at positions 5322 through 5461). To facilitate comparison with the STM data, the guanine sites are indicated by red characters and are also connected by red arrows to the corresponding parts of the image. (Reprinted with permission from Nature Publishing Group) | |
| Currently, there are two main problems with this STM technique: One, to become viable as a practical tool it must be able to clearly recognize all four types of base molecule of DNA, not just guanine. Two, and a killer as far as commercial or large-scale applications are concerned, is the time it takes to perform the sequencing. It took almost an hour to obtain the dI/dV map of image d in the above figure. The researchers point out that, in general, lock-in detection sacrifices temporal resolution for the sake of improved signal-to-noise ratio. "In our measurement system, to obtain the dI/dV spectrum, it was necessary to stop the scanning and feedback of the STM probe and then measure the I?V characteristic, so that a time ranging from 1 second to 1 minute was needed for each point." | |
| Kawai and Tanaka explain that if STM software and control mechanisms capable of finding chain-shaped polymers such as DNA can be developed, then the time taken to scan parts where the sample is not present can be greatly reduced. "Savings in cost and time can also be made if sequencing is performed from the topographic image alone without using a lock-in amplifier, but it would still be necessary to use a method for identifying contamination." | |
 By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
|
|
|
Become a Spotlight guest author! Join our large and growing group of guest contributors. Have you just published a scientific paper or have other exciting developments to share with the nanotechnology community? Here is how to publish on nanowerk.com. |
|
