Showing Spotlights 25 - 32 of 68 in category All (newest first):
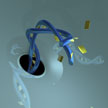 DNA is constantly being damaged in our cells by radiation and other random sources. One of the major forms of this damage is called depurination, or the selective loss of A and G bases from the double helix structure. In our cells, there is a system in place to fix depurination. It usually is quite successful at repairing the damage, but can sometimes make mistakes that result in mutations. As a result, depurination is directly linked to a host of diseases, including anemia and cancer. In new work, researchers show that DNA depurination can be detected electrically using solid-state nanopores.
DNA is constantly being damaged in our cells by radiation and other random sources. One of the major forms of this damage is called depurination, or the selective loss of A and G bases from the double helix structure. In our cells, there is a system in place to fix depurination. It usually is quite successful at repairing the damage, but can sometimes make mistakes that result in mutations. As a result, depurination is directly linked to a host of diseases, including anemia and cancer. In new work, researchers show that DNA depurination can be detected electrically using solid-state nanopores.
Aug 7th, 2014
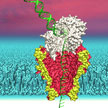 Sequencing technologies have made it cheaper and faster to read the sequence of bases on a strand of DNA. A promising technology to take these advances further is nanopore sequencing. Individual strands of DNA are moved through a nanopore gap not much wider than the DNA itself. As the DNA passes through the nanopore, continuous information is gained about the sequence of individual bases - the A, C, G and Ts that make up DNA. Researchers now have developed a nanopore sequencing technique reaching read lengths of several thousand bases.
Sequencing technologies have made it cheaper and faster to read the sequence of bases on a strand of DNA. A promising technology to take these advances further is nanopore sequencing. Individual strands of DNA are moved through a nanopore gap not much wider than the DNA itself. As the DNA passes through the nanopore, continuous information is gained about the sequence of individual bases - the A, C, G and Ts that make up DNA. Researchers now have developed a nanopore sequencing technique reaching read lengths of several thousand bases.
Jul 17th, 2014
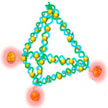 One way to eliminate the toxicity issue of synthetic nanomaterials used in nanomedicine is by working with truly biocompatible natural carriers for sensing and drug delivery applications. The emerging field of DNA nanotechnology may provide a solution. In new work, researchers have developed a novel theranostic platform which is made by utilizing a self-assembled DNA nanopyramid as scaffold for incorporation of both detection and therapeutic moieties to combat bacterial infection.
One way to eliminate the toxicity issue of synthetic nanomaterials used in nanomedicine is by working with truly biocompatible natural carriers for sensing and drug delivery applications. The emerging field of DNA nanotechnology may provide a solution. In new work, researchers have developed a novel theranostic platform which is made by utilizing a self-assembled DNA nanopyramid as scaffold for incorporation of both detection and therapeutic moieties to combat bacterial infection.
Jul 11th, 2014
 Nanomaterials for nanomedicine and biological applications are often two-component structures - referred to as 'nanoconstructs' -consisting of a 'hard' nanoparticle core and a 'soft' shell of biomolecular ligands. Researchers have now demonstrated a nanoconstruct with enhanced in vitro efficacy. This highly loaded nanoconstruct was taken up by pancreatic cancer cells and fibrosarcoma cells at fast rates. The team found that the increased loading of Apt on AuNS also resulted in an enhanced in vitro response.
Nanomaterials for nanomedicine and biological applications are often two-component structures - referred to as 'nanoconstructs' -consisting of a 'hard' nanoparticle core and a 'soft' shell of biomolecular ligands. Researchers have now demonstrated a nanoconstruct with enhanced in vitro efficacy. This highly loaded nanoconstruct was taken up by pancreatic cancer cells and fibrosarcoma cells at fast rates. The team found that the increased loading of Apt on AuNS also resulted in an enhanced in vitro response.
Apr 9th, 2014
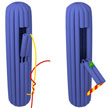 Not to be confused with the nanorobots of science fiction, for medical nanotechnology researchers a nanorobot, or nanobot, is a popular term for molecules with a unique property that enables them to be programmed to carry out a specific task. In what is the smallest 3D DNA origami box so far, researchers in Italy have now fabricated a nanorobot with a switchable flap that, when instructed with a freely defined molecular message, can perform a specifically programmed duty. Slightly larger nanocontainers with a controllable lid have already been demonstrated by others to be suitable for the delivery of drugs or molecular signals, but this new cylindrical nanobot has an innovative opening mechanism.
Not to be confused with the nanorobots of science fiction, for medical nanotechnology researchers a nanorobot, or nanobot, is a popular term for molecules with a unique property that enables them to be programmed to carry out a specific task. In what is the smallest 3D DNA origami box so far, researchers in Italy have now fabricated a nanorobot with a switchable flap that, when instructed with a freely defined molecular message, can perform a specifically programmed duty. Slightly larger nanocontainers with a controllable lid have already been demonstrated by others to be suitable for the delivery of drugs or molecular signals, but this new cylindrical nanobot has an innovative opening mechanism.
Apr 2nd, 2014
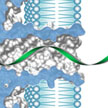 Epigenetic mechanisms are chemical changes in DNA that do not alter the actual genetic code but can influence the expression of genes, and can be passed on when cells reproduce. One of the most important is DNA methylation, where methyl groups - small structures of carbon and hydrogen - are appended to specific locations on a DNA strand. Recently, both biological and synthetic nanopores have been proposed for DNA methylation detection. In new work, researchers employed protein nanopores to investigate a novel metal ion-bridged DNA interstrand lock, and explore its potential in location-specific methylation detection.
Epigenetic mechanisms are chemical changes in DNA that do not alter the actual genetic code but can influence the expression of genes, and can be passed on when cells reproduce. One of the most important is DNA methylation, where methyl groups - small structures of carbon and hydrogen - are appended to specific locations on a DNA strand. Recently, both biological and synthetic nanopores have been proposed for DNA methylation detection. In new work, researchers employed protein nanopores to investigate a novel metal ion-bridged DNA interstrand lock, and explore its potential in location-specific methylation detection.
Oct 21st, 2013
 DNA is a powerful biomaterial for creating rationally designed and functionally enhanced nanostructures. Emerging DNA nanotechnology employs DNA as a programmable building material for self-assembled, nanoscale structures. Researchers have also shown that DNA nanotechnology can be integrated with traditional silicon processing. DNA nanoarchitectures positioned at substrate interfaces can offer unique advantages leading to improved surface properties relevant to biosensing (for instance, graphene and DNA can combine to create a stable and accurate biosensor), nanotechnology, materials science, and cell biology.
DNA is a powerful biomaterial for creating rationally designed and functionally enhanced nanostructures. Emerging DNA nanotechnology employs DNA as a programmable building material for self-assembled, nanoscale structures. Researchers have also shown that DNA nanotechnology can be integrated with traditional silicon processing. DNA nanoarchitectures positioned at substrate interfaces can offer unique advantages leading to improved surface properties relevant to biosensing (for instance, graphene and DNA can combine to create a stable and accurate biosensor), nanotechnology, materials science, and cell biology.
Feb 7th, 2013
 Recent developments in nanotechnology have enabled significant improvement in the field of anti-counterfeiting measures. One company for instance is working on fluorescent nanostructures to improve banknote security; another one has developed DNA tags for deposition on nanoelectronics wafers and computer chips to ensure the integrity and security of processed wafers. DNA-based protection technologies are especially suitable for anti-counterfeiting measures.The DNA molecules are added a products raw material during the production process. Only 1 ppm (one part per million) is required to uniquely mark the material The DNA molecular structure can then be read as a mathematical code based on the four DNA molecules. So a DNA code, in contrast to the binary code used in IT security, is a combination of the letters A, C, G and T. A 10-digit code could look like this: C-G-A-C-T-T-G-A-C-A.
Recent developments in nanotechnology have enabled significant improvement in the field of anti-counterfeiting measures. One company for instance is working on fluorescent nanostructures to improve banknote security; another one has developed DNA tags for deposition on nanoelectronics wafers and computer chips to ensure the integrity and security of processed wafers. DNA-based protection technologies are especially suitable for anti-counterfeiting measures.The DNA molecules are added a products raw material during the production process. Only 1 ppm (one part per million) is required to uniquely mark the material The DNA molecular structure can then be read as a mathematical code based on the four DNA molecules. So a DNA code, in contrast to the binary code used in IT security, is a combination of the letters A, C, G and T. A 10-digit code could look like this: C-G-A-C-T-T-G-A-C-A.
Nov 6th, 2012
 DNA is constantly being damaged in our cells by radiation and other random sources. One of the major forms of this damage is called depurination, or the selective loss of A and G bases from the double helix structure. In our cells, there is a system in place to fix depurination. It usually is quite successful at repairing the damage, but can sometimes make mistakes that result in mutations. As a result, depurination is directly linked to a host of diseases, including anemia and cancer. In new work, researchers show that DNA depurination can be detected electrically using solid-state nanopores.
DNA is constantly being damaged in our cells by radiation and other random sources. One of the major forms of this damage is called depurination, or the selective loss of A and G bases from the double helix structure. In our cells, there is a system in place to fix depurination. It usually is quite successful at repairing the damage, but can sometimes make mistakes that result in mutations. As a result, depurination is directly linked to a host of diseases, including anemia and cancer. In new work, researchers show that DNA depurination can be detected electrically using solid-state nanopores.
 Subscribe to our Nanotechnology Spotlight feed
Subscribe to our Nanotechnology Spotlight feed





