Showing Spotlights 33 - 40 of 68 in category All (newest first):
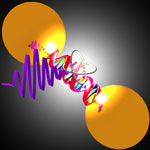 The emission of light by a single molecule is a cornerstone of nano-optics that will enable applications in quantum information processing or single-molecule spectroscopy. However, a key challenge in nano-optics is to bring light to and collect light from nano-scale systems. In conventional electronics, the interconnect between locally stored and radiated signals, for example radio broadcasts or mobile phone transmissions, is formed by antennas. For an antenna to work at the wavelength of light it is necessary to downscale the structure by the same factor as the wavelength or the frequency of the wave, i.e. roughly by a factor of 10 million. Once the nanofabrication issues are sorted out, nano-optical antennas could become ubiquitous in all applications based on light-matter interactions such as sensing, light emission (e.g. LEDs) and detection, as well as light harvesting, i.e. for solar cell applications.
The emission of light by a single molecule is a cornerstone of nano-optics that will enable applications in quantum information processing or single-molecule spectroscopy. However, a key challenge in nano-optics is to bring light to and collect light from nano-scale systems. In conventional electronics, the interconnect between locally stored and radiated signals, for example radio broadcasts or mobile phone transmissions, is formed by antennas. For an antenna to work at the wavelength of light it is necessary to downscale the structure by the same factor as the wavelength or the frequency of the wave, i.e. roughly by a factor of 10 million. Once the nanofabrication issues are sorted out, nano-optical antennas could become ubiquitous in all applications based on light-matter interactions such as sensing, light emission (e.g. LEDs) and detection, as well as light harvesting, i.e. for solar cell applications.
Jul 31st, 2012
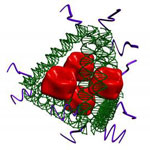 Vaccination is one of the most effective ways to prevent microbial infection. Synthetic vaccines can combine a portion of a microbe, known as an 'antigen' together with an adjuvant that stimulates the immune system. Delivering both the adjuvant and antigen to the appropriate immune cells is challenging. DNA nanotechnology may provide a solution by acting as a scaffold to co-deliver both antigen and adjuvant. However, the potential of DNA nanostructure-based vaccines has only been demonstrated in vitro. Now, a team of researchers based out of Arizona State University demonstrated that DNA nanostructures with appended adjuvants could elicit antibody production against a model antigen in mice.
Vaccination is one of the most effective ways to prevent microbial infection. Synthetic vaccines can combine a portion of a microbe, known as an 'antigen' together with an adjuvant that stimulates the immune system. Delivering both the adjuvant and antigen to the appropriate immune cells is challenging. DNA nanotechnology may provide a solution by acting as a scaffold to co-deliver both antigen and adjuvant. However, the potential of DNA nanostructure-based vaccines has only been demonstrated in vitro. Now, a team of researchers based out of Arizona State University demonstrated that DNA nanostructures with appended adjuvants could elicit antibody production against a model antigen in mice.
Jul 30th, 2012
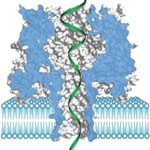 Every aspect of cellular activities, including cell proliferation, differentiation, metabolism and apoptosis, can be regulated by a class of tiny but very important nucleic acids fragments called microRNAs (miRNAs). They bind to specific messenger RNAs and cause messenger RNA degradation or inhibit translation, thereby regulate gene expression at the post-translational level. In cancer cells, the homeostasis of these normal biological processes is disrupted, partially by dysregulated miRNAs, therefore the level of microRNAs is an indicator to the disease development, and miRNAs in cancer tissues or biofluids can be utilized as a diagnostic biomarker for cancer detection. Now, researchers report a miRNAs-based discovery that could provide a much earlier warning signal for lung cancer.
Every aspect of cellular activities, including cell proliferation, differentiation, metabolism and apoptosis, can be regulated by a class of tiny but very important nucleic acids fragments called microRNAs (miRNAs). They bind to specific messenger RNAs and cause messenger RNA degradation or inhibit translation, thereby regulate gene expression at the post-translational level. In cancer cells, the homeostasis of these normal biological processes is disrupted, partially by dysregulated miRNAs, therefore the level of microRNAs is an indicator to the disease development, and miRNAs in cancer tissues or biofluids can be utilized as a diagnostic biomarker for cancer detection. Now, researchers report a miRNAs-based discovery that could provide a much earlier warning signal for lung cancer.
Sep 20th, 2011
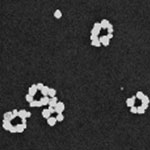 DNA origami is a design technique that is used by nanotechnology researchers to fold DNA strands into something resembling a programmable pegboard on which different nanocomponents can be attached. These DNA assemblies allow the bottom-up fabrication of complex nanostructures with arbitrary shapes and patterns on a 100 nm scale. For instance, DNA origami have been heralded as a potential breakthrough for the creation of nanoscale circuits and devices. DNA can also be metallized with different metals, resulting in conducting nanowires. Researchers have now have developed a method to assemble metallic nanocircuits with arbitrary shapes, by attaching metallic nanoparticles to select locations of the DNA origami and then fusing them to form wires, rings, or any other complex shape. These pre-designed structures are programmed by fully utilizing the self-assembling and recognition properties of DNA.
DNA origami is a design technique that is used by nanotechnology researchers to fold DNA strands into something resembling a programmable pegboard on which different nanocomponents can be attached. These DNA assemblies allow the bottom-up fabrication of complex nanostructures with arbitrary shapes and patterns on a 100 nm scale. For instance, DNA origami have been heralded as a potential breakthrough for the creation of nanoscale circuits and devices. DNA can also be metallized with different metals, resulting in conducting nanowires. Researchers have now have developed a method to assemble metallic nanocircuits with arbitrary shapes, by attaching metallic nanoparticles to select locations of the DNA origami and then fusing them to form wires, rings, or any other complex shape. These pre-designed structures are programmed by fully utilizing the self-assembling and recognition properties of DNA.
Aug 11th, 2011
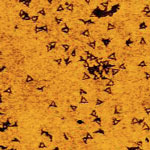 To build microprocessors with more than one billion transistors, manufacturers still use the same technique - photolithography, the high-tech, nanoscale version of printing technology - that they have been using for the past 50 years. State-of-the-art photolithography processes use 193 nm light to produce diffraction-limited features as small as 32 nm. Going beyond 32 nm, the cost and complexity rises significantly, posing massive technological and economic challenges for chip manufacturers. This provides plenty of incentives for researchers to explore alternative manufacturing technologies for chipmakers. One novel approach is based on the use of DNA nanostructures to pattern a silicon wafer.
To build microprocessors with more than one billion transistors, manufacturers still use the same technique - photolithography, the high-tech, nanoscale version of printing technology - that they have been using for the past 50 years. State-of-the-art photolithography processes use 193 nm light to produce diffraction-limited features as small as 32 nm. Going beyond 32 nm, the cost and complexity rises significantly, posing massive technological and economic challenges for chip manufacturers. This provides plenty of incentives for researchers to explore alternative manufacturing technologies for chipmakers. One novel approach is based on the use of DNA nanostructures to pattern a silicon wafer.
Jul 26th, 2011
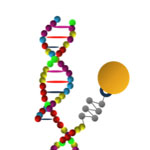 MicroRNAs (miRNAs) are short ribonucleic acid molecules, consisting of 21-25 nucleotide bases, that negatively regulate gene expression, also termed as gene silencing. Each miRNA is thought to regulate multiple genes, and since the human genome encodes hundreds of miRNAs, the potential regulatory circuitry afforded by miRNA is enormous. Recent discoveries suggest the association of specific miRNA sequences with a spectrum of diseases including cardiovascular and autoimmune diseases, as well as with a variety of cancers. It is therefore imperative, for diagnostics and prognostics, to accurately measure the expression levels of target miRNA molecules in patients' tissue samples or body fluids. To that end, researchers have developed an alternative way for the direct analysis of miRNAs in an array format, demonstrating fast and ultrasensitive detection of specific miRNAs.
MicroRNAs (miRNAs) are short ribonucleic acid molecules, consisting of 21-25 nucleotide bases, that negatively regulate gene expression, also termed as gene silencing. Each miRNA is thought to regulate multiple genes, and since the human genome encodes hundreds of miRNAs, the potential regulatory circuitry afforded by miRNA is enormous. Recent discoveries suggest the association of specific miRNA sequences with a spectrum of diseases including cardiovascular and autoimmune diseases, as well as with a variety of cancers. It is therefore imperative, for diagnostics and prognostics, to accurately measure the expression levels of target miRNA molecules in patients' tissue samples or body fluids. To that end, researchers have developed an alternative way for the direct analysis of miRNAs in an array format, demonstrating fast and ultrasensitive detection of specific miRNAs.
Jun 6th, 2011
 A Single Nucleotide Polymorphism (SNP) is a single nucleotide replacement in a DNA sequence - occurring when a single nucleotide (A, T, C, or G) in the genome differs - which can result in different reaction by people to pathogens and medicines. Detection of these SNPs is becoming increasingly important with the move towards more personalized healthcare. Researchers are therefore working hard in developing biomedical lab-on-chip sensors that allow the fast detection of SNPs in DNA using only very small samples of a patient's blood. Already, nanoscale detection techniques such as synthetic nanochannels are being used for DNA detection by specific DNA hybridization with molecular probes immobilized on the nanochannel walls. However, the preparation of these sensors is not easy and specific functionalization at the wall surface remains a critical issues. Researchers have now introduced a new concept of DNA-based molecular recognition agents which allows detecting SNPs with very high precision and efficiency.
A Single Nucleotide Polymorphism (SNP) is a single nucleotide replacement in a DNA sequence - occurring when a single nucleotide (A, T, C, or G) in the genome differs - which can result in different reaction by people to pathogens and medicines. Detection of these SNPs is becoming increasingly important with the move towards more personalized healthcare. Researchers are therefore working hard in developing biomedical lab-on-chip sensors that allow the fast detection of SNPs in DNA using only very small samples of a patient's blood. Already, nanoscale detection techniques such as synthetic nanochannels are being used for DNA detection by specific DNA hybridization with molecular probes immobilized on the nanochannel walls. However, the preparation of these sensors is not easy and specific functionalization at the wall surface remains a critical issues. Researchers have now introduced a new concept of DNA-based molecular recognition agents which allows detecting SNPs with very high precision and efficiency.
Feb 25th, 2011
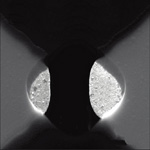 Researchers worldwide are working on fast and low-cost strategies to sequence DNA, that is, to read off the content of our genome. Particularly promising for future genome sequencing are devices that measure single molecules. In this respect, the creation of nanochannels or nanopores in thin membranes has attracted much interest due to the potential to isolate and sense single DNA molecules while they translocate through the highly confined channels. Particularly interesting are techniques that can offer fast and low cost readout of long DNA molecules without the need of DNA labelling or amplification. In very interesting work performed at Imperial College London, researchers have now successfully developed a protocol for the fabrication of a solid state nanopore aligned to a tunneling junction.
Researchers worldwide are working on fast and low-cost strategies to sequence DNA, that is, to read off the content of our genome. Particularly promising for future genome sequencing are devices that measure single molecules. In this respect, the creation of nanochannels or nanopores in thin membranes has attracted much interest due to the potential to isolate and sense single DNA molecules while they translocate through the highly confined channels. Particularly interesting are techniques that can offer fast and low cost readout of long DNA molecules without the need of DNA labelling or amplification. In very interesting work performed at Imperial College London, researchers have now successfully developed a protocol for the fabrication of a solid state nanopore aligned to a tunneling junction.
Dec 13th, 2010
 The emission of light by a single molecule is a cornerstone of nano-optics that will enable applications in quantum information processing or single-molecule spectroscopy. However, a key challenge in nano-optics is to bring light to and collect light from nano-scale systems. In conventional electronics, the interconnect between locally stored and radiated signals, for example radio broadcasts or mobile phone transmissions, is formed by antennas. For an antenna to work at the wavelength of light it is necessary to downscale the structure by the same factor as the wavelength or the frequency of the wave, i.e. roughly by a factor of 10 million. Once the nanofabrication issues are sorted out, nano-optical antennas could become ubiquitous in all applications based on light-matter interactions such as sensing, light emission (e.g. LEDs) and detection, as well as light harvesting, i.e. for solar cell applications.
The emission of light by a single molecule is a cornerstone of nano-optics that will enable applications in quantum information processing or single-molecule spectroscopy. However, a key challenge in nano-optics is to bring light to and collect light from nano-scale systems. In conventional electronics, the interconnect between locally stored and radiated signals, for example radio broadcasts or mobile phone transmissions, is formed by antennas. For an antenna to work at the wavelength of light it is necessary to downscale the structure by the same factor as the wavelength or the frequency of the wave, i.e. roughly by a factor of 10 million. Once the nanofabrication issues are sorted out, nano-optical antennas could become ubiquitous in all applications based on light-matter interactions such as sensing, light emission (e.g. LEDs) and detection, as well as light harvesting, i.e. for solar cell applications.
 Subscribe to our Nanotechnology Spotlight feed
Subscribe to our Nanotechnology Spotlight feed





