Showing Spotlights 9 - 16 of 110 in category All (newest first):
 Atomic force microscopy (AFM) is one of the newer techniques available for virus research. AFM is a cantilever-based technique that utilizes a sharp tip to interrogate surfaces at resolutions well below the optical diffraction limit. Beyond imaging, AFM is also a powerful tool for nano-mechanical probing and nano-manipulation. One of the primary advantages of AFM is that it can operate on samples immersed in liquid. This empowers experiments on living cells at physiologically relevant conditions.
Atomic force microscopy (AFM) is one of the newer techniques available for virus research. AFM is a cantilever-based technique that utilizes a sharp tip to interrogate surfaces at resolutions well below the optical diffraction limit. Beyond imaging, AFM is also a powerful tool for nano-mechanical probing and nano-manipulation. One of the primary advantages of AFM is that it can operate on samples immersed in liquid. This empowers experiments on living cells at physiologically relevant conditions.
Oct 23rd, 2020
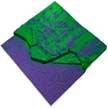 Kelvin probe force microscopy (KPFM), also known as surface potential microscopy, is one member of a suite of electrical characterization methods available in atomic force microscopes. It maps the contact potential difference between a surface and the cantilever, containing information about the surface potential and work function. KPFM is a surface-sensitive method that probes at and near the surface only. It is often used as a qualitative technique to obtain contrast based on the surface potential.
Kelvin probe force microscopy (KPFM), also known as surface potential microscopy, is one member of a suite of electrical characterization methods available in atomic force microscopes. It maps the contact potential difference between a surface and the cantilever, containing information about the surface potential and work function. KPFM is a surface-sensitive method that probes at and near the surface only. It is often used as a qualitative technique to obtain contrast based on the surface potential.
Oct 21st, 2020
 The foremost atom of the tip in a scanning probe microscope is critically important for precise imaging results. In atomic force microscopy, the front atom significantly affects atomic-scale contrast and atom manipulation. The chemical species is a dominant factor in determining the tip state, and in situ chemical identification of a tip apex remains a challenging task. Researchers found that Pauling's equation is applicable to the analysis of tip apex atoms. The team also demonstrated a way to determine electronegativity of surface atoms solely by experiments.
The foremost atom of the tip in a scanning probe microscope is critically important for precise imaging results. In atomic force microscopy, the front atom significantly affects atomic-scale contrast and atom manipulation. The chemical species is a dominant factor in determining the tip state, and in situ chemical identification of a tip apex remains a challenging task. Researchers found that Pauling's equation is applicable to the analysis of tip apex atoms. The team also demonstrated a way to determine electronegativity of surface atoms solely by experiments.
Feb 21st, 2020
 Two-dimensional (2D) materials could offer new building blocks for future technologies, but this requires approaches to control the carrier type in 2D semiconductors. A number of pioneering works have demonstrated different methods to program the carrier type in 2D materials, such as electrostatic doping, chemical doping, ion implantation, charge transfer, and annealing control. Recently, a team of researchers have developed a technique to dope 2D materials for redefinable nanoelectronics using nonvolatile ferroelectric domains.
Two-dimensional (2D) materials could offer new building blocks for future technologies, but this requires approaches to control the carrier type in 2D semiconductors. A number of pioneering works have demonstrated different methods to program the carrier type in 2D materials, such as electrostatic doping, chemical doping, ion implantation, charge transfer, and annealing control. Recently, a team of researchers have developed a technique to dope 2D materials for redefinable nanoelectronics using nonvolatile ferroelectric domains.
Feb 20th, 2020
 Nanoparticles have shown a lot of promise in biomedical applications. However, accumulation of nanoparticles in the liver is a major concern, and may be one of the greatest barriers to the widespread adoption of nanoparticles in the clinic. This is especially true for metallic nanoparticles, since the long-term effects of their accumulation in the liver has not been widely studied.
A team of researchers has looked to nature for inspiration in solving this problem. They decided to use biliverdin, a bile pigment, as the building block for their nanoparticles.
Nanoparticles have shown a lot of promise in biomedical applications. However, accumulation of nanoparticles in the liver is a major concern, and may be one of the greatest barriers to the widespread adoption of nanoparticles in the clinic. This is especially true for metallic nanoparticles, since the long-term effects of their accumulation in the liver has not been widely studied.
A team of researchers has looked to nature for inspiration in solving this problem. They decided to use biliverdin, a bile pigment, as the building block for their nanoparticles.
Nov 18th, 2019
 By precisely stacking various 2D materials in a predetermined sequence on top of each other, researchers create van der Waals heterostructures that, due to their unique interlayer coupling, have special optoelectronic properties. Unfortunately, the study of 2D monolayers is plagued by trapped contaminants in between the 2D sheets as well as between the 2D sheets and the underlying substrate. These contaminants make it difficult to obtain precise and reproducible experimental observations. A simple technique for removing these contaminants in a process similar to a squeegee.
By precisely stacking various 2D materials in a predetermined sequence on top of each other, researchers create van der Waals heterostructures that, due to their unique interlayer coupling, have special optoelectronic properties. Unfortunately, the study of 2D monolayers is plagued by trapped contaminants in between the 2D sheets as well as between the 2D sheets and the underlying substrate. These contaminants make it difficult to obtain precise and reproducible experimental observations. A simple technique for removing these contaminants in a process similar to a squeegee.
May 1st, 2018
 Among the important parameters in optical lithography is the spatial resolution you can get and the time you need to draw your pattern. Systems with regular and fixed patterns can be extremely fast but those systems are based on masks. Mask production is a time consuming and expensive process. So-called mask-less systems can draw unorganized patterns directly on substrates, at the cost of longer process times. Researchers now have presented a new lithographic approach with a high-resolution, low-cost technique based on nanosphere lithography.
Among the important parameters in optical lithography is the spatial resolution you can get and the time you need to draw your pattern. Systems with regular and fixed patterns can be extremely fast but those systems are based on masks. Mask production is a time consuming and expensive process. So-called mask-less systems can draw unorganized patterns directly on substrates, at the cost of longer process times. Researchers now have presented a new lithographic approach with a high-resolution, low-cost technique based on nanosphere lithography.
Oct 4th, 2017
 Photothermal induced resonance (PTIR) has found application in the characterization of materials in fields spanning from photovoltaics, plasmonic, polymer science, biology and geology to name a few. PTIR combines the spatial resolution of atomic force microscopy with the specificity of absorption spectroscopy, enabling mapping of composition and electronic bandgap, material identification and biomolecule conformational analysis with nanoscale spatial resolution. Scientists have now implemented, for the first time, an integrated near-field cavity-optomechanics readout concept to realize fully functional nanoscale AFM probes capable of ultralow detection noise within an extremely wide measurement bandwidth in ambient conditions, surpassing all previous AFM probes.
Photothermal induced resonance (PTIR) has found application in the characterization of materials in fields spanning from photovoltaics, plasmonic, polymer science, biology and geology to name a few. PTIR combines the spatial resolution of atomic force microscopy with the specificity of absorption spectroscopy, enabling mapping of composition and electronic bandgap, material identification and biomolecule conformational analysis with nanoscale spatial resolution. Scientists have now implemented, for the first time, an integrated near-field cavity-optomechanics readout concept to realize fully functional nanoscale AFM probes capable of ultralow detection noise within an extremely wide measurement bandwidth in ambient conditions, surpassing all previous AFM probes.
Sep 1st, 2017
 Atomic force microscopy (AFM) is one of the newer techniques available for virus research. AFM is a cantilever-based technique that utilizes a sharp tip to interrogate surfaces at resolutions well below the optical diffraction limit. Beyond imaging, AFM is also a powerful tool for nano-mechanical probing and nano-manipulation. One of the primary advantages of AFM is that it can operate on samples immersed in liquid. This empowers experiments on living cells at physiologically relevant conditions.
Atomic force microscopy (AFM) is one of the newer techniques available for virus research. AFM is a cantilever-based technique that utilizes a sharp tip to interrogate surfaces at resolutions well below the optical diffraction limit. Beyond imaging, AFM is also a powerful tool for nano-mechanical probing and nano-manipulation. One of the primary advantages of AFM is that it can operate on samples immersed in liquid. This empowers experiments on living cells at physiologically relevant conditions.
 Subscribe to our Nanotechnology Spotlight feed
Subscribe to our Nanotechnology Spotlight feed





