| Posted: Nov 29, 2012 | |
Injecting femtotliter quantities into single cell nuclei using fluidic force microscopy (w/video) |
|
| (Nanowerk Spotlight) Fluidic force microscopy (FluidFM) is an emerging technology which combines atomic force microscopy (AFM) with microfluidics (described in detail in "FluidFM: Combining Atomic Force Microscopy and Nanofluidics in a Universal Liquid Delivery System for Single Cell Applications and Beyond"). | |
| In a new study, researchers in Switzerland have now developed an innovative method for straightforward injection into the nucleus of a living cell, taking advantage of the nanoscale accuracy and small probe size of AFM and the possibility to handle fluid under pressure-control through the integrated microchannel. | |
| "Invented in 1904 and optimized in the 1970s, microinjection using glass pipettes represents the most widely used approach for intracellular injection," Julia A. Vorholt, a professor at the Institute of Microbiology at ETH Zurich, explains to Nanowerk. "However, microinjection has major limitations, such as the manual dexterity required to handle glass micropipettes, the relatively high risk to damage the cells, the difficulties to deliver small (femtoliter) volumes, and the lack of readout to monitor the penetration of the pipette through the cell membrane(s)." | |
| To overcome these limitations, Vorholt and her group, working with scientists from ETH's Laboratory of Biosensors and Bioelectronics, developed a protocol that makes the intranuclear injection a precise and reliable procedure. | |
 |
|
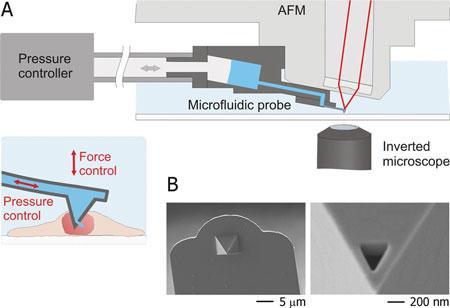
| A) Schematic representation of the intranuclear injection using fluidic force microscopy. The setup features an inverted microscope, an AFM, and a microfluidic probe connected to a pressure controller. Optical-, force- and pressure-monitoring are performed simultaneously along the entire intranuclear injection process. B) Scanning electron micrographs of a FluidFM injection probe; a triangular aperture was milled by focused ion beam at the front facet of the pyramidal tip. (Reprinted with permission from Wiley-VCH Verlag) | |
| Reporting their findings in the November 20, 2012 online edition of Small ("Force-Controlled Fluidic Injection into Single Cell Nuclei"), first-authored by Orane Guillaume-Gentil, a post-doc in Vorholt's group, the team describes its design for an optimized force spectroscopy routine to reliably insert, maintain, and withdraw the microfluidic probe into/from single cell nuclei. | |
| In addition, continuous force-monitoring enabled the detection in situ of the probe penetration through the cytoplasmic and nuclear membranes. Once the probe inserted in the intranuclear compartment, a defined pressure pulse was applied to release the biomolecule solution. | |
| "By finely tuning the pressure pulses, we could inject volumes ranging from one or two picoliters down to a few tens of femtoliters" says Guillaume-Gentil. | |
| The team demonstrated the high potential of this approach in single-cell transfection experiments in which single-cell expression of green fluorescent protein (GFP) was induced by the intranuclear injection of plasmid DNA encoding the fluorescent protein. | |
| "Our approach uniquely enabled reliable probe insertion with a simultaneous readout of the membrane perforation events, maximal injection efficiency (100%), minimal damage to the cell – cells survived injection, expressed proteins encoded in an injected plasmid, and divided as in standard culture conditions – and control over the volumes injected," Vorholt summarizes the team's results. | |
 |
|
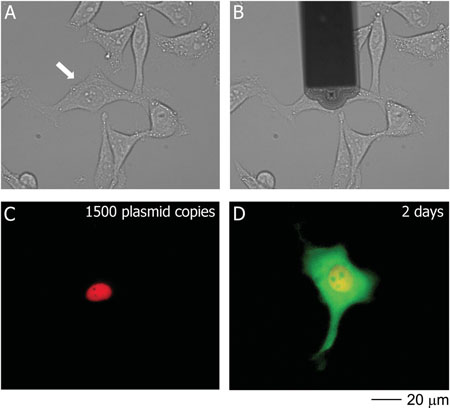
| Transfection of a HeLa cell with pmaxGFP; the phase contrast images show the cell before (A) and during (B) injection. The fluorescence images show the cell immediately (C) and 2 days (D) after injection. DexTRITC (red) co-injected with the plasmid remained localized in the nucleus, and 2 days post-injection the cell was producing GFP (green). The arrow in A) indicates the targeted cell. (Reprinted with permission from Wiley-VCH Verlag) | |
| The potential applications for straightforward and standardized injection into cell nuclei with FluidFM are wide-ranging. Vorholt points out that the possibility to release defined quantities of DNA, RNA, or proteins directly into single cell nucleus allows for gaining new insights into critical intracellular processes at the single-cell level. "The technology will enable examining the role of specific genes or proteins, offering great opportunities in various field of cell biology research such as neuroscience, cancer research, or stem cell studies,". | |
| She also points out that, besides intranuclear injection, fluidic force microscopy has shown great promises for a broad range of applications in particular in biology: cytosolic injection ("FluidFM: Combining Atomic Force Microscopy and Nanofluidics in a Universal Liquid Delivery System for Single Cell Applications and Beyond"), spatial manipulation of virus, bacteria, yeasts, and mammalian cells ("Force-controlled spatial manipulation of viable mammalian cells and micro-organisms by means of FluidFM technology"), dynamic and serial adhesion measurements of single yeast and mammalian cells (Potthoff et al., PLoS ONE 2012, in press). | |
| The technology is currently further developed, expanding its high versatility with additional applications such as cell sorting, patch-clamping, biosensing, extraction, and nanolithography. | |
 By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
|
|
|
Become a Spotlight guest author! Join our large and growing group of guest contributors. Have you just published a scientific paper or have other exciting developments to share with the nanotechnology community? Here is how to publish on nanowerk.com. |
|
