| Posted: Oct 14, 2013 | |
Robust 3D atomic force microscopy without the need for lateral scanning |
|
| (Nanowerk Spotlight) The realization of a three-dimensional tomic force microscopy (3D AFM) portends exciting research directions across nanoscience and nanotechnology. Demonstrations to date have been limited by the indirect means that are required to extract a three-dimensional force vector from the traditional 1D observable in AFM (i.e., cantilever deflection). | |
| Existing 3D AFM techniques require recording thousands of frequency shift curves at different lateral locations followed by off-line integration (to yield energy) and lateral differentiation (to yield lateral force). This procedure is inherently slow. Thus, it is largely restricted to studies of static samples. Best results to date in terms of processing time (53 seconds for a 3D force map) have come from a group at the University of Kanazawa in Japan, but their method still relies on the traditional one-dimensional AFM observable and requires lateral scanning (see: "Three-dimensional quantitative force maps in liquid with 10 piconewton, angstrom and sub-minute resolutions"). | |
| "In most imaging and spectroscopy applications, force measurements are made in the z direction, that is, normal to the sample surface," Gavin M. King, an Assistant Professor in the Department of Physics and Astronomy at the University of Missouri-Columbia, explains to Nanowerk. "However, torsional deflection – twisting – of the cantilever can also be monitored, as in frictional force microscopy. In both cases, tip motion, which occurs in three dimensions, is convolved into angular displacements of the cantilever. It is challenging to infer 3D tip trajectories from this reduced coordinate system, even if the geometry of the tip and its orientation relative to the sample are known. This is because tip and cantilever dynamics are not always in lock-step with each other as transient excitations can propagate along the flexible cantilever, especially during fast scanning." | |
| In the October 7, 2013, online edition of Nano Letters ("Three-Dimensional Atomic Force Microscopy: Interaction Force Vector by Direct Observation of Tip Trajectory"), King and his group now report 3D force measurements based on a 3D local observable, rather than on cantilever deflection alone. | |
| An overarching challenge in performing 3D AFM in biological fluid is to measure small forces in a perturbative environment rapidly and with high precision. This paper takes a step towards direction. The central thesis that they discuss and demonstrate in it is that a 3D AFM should have a 3D observable at its core. | |
 |
|
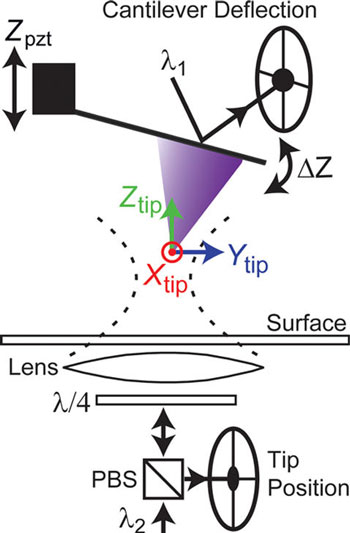
| Schematic of the apparatus. The combination of 3D tip position measurement (Xtip, Ytip, Ztip) with conventional force detection is shown. An optical lever arm laser (λ1) is reflected off the back of the cantilever to yield ΔZ. A separate laser (λ2) is focused onto the tip apex using an objective lens positioned under the sample. Backscattered light from the tip is collected by the same lens and detected using a quadrant photo detector, yielding the three-dimensional tip position. (Reprinted with permission from American Chemical Society) | |
| The team's measurements build upon ultrastable AFM, a recently developed technique that was inspired by surface-coupled optical trapping microscopy methods (see: "Ultrastable Atomic Force Microscopy: Atomic-Scale Stability and Registration in Ambient Conditions"). | |
| "Ultrastable AFM employs a focused laser that backscatters off the tip itself to rapidly yield tip position with high spatial precision in 3D," King describes the technique. "In our previous work, this tip-position data was used to stabilize the tip with respect to a surface in contact mode and in air. Here, we extend that work to the commonly applied intermittent contact (tapping) mode in fluid and use the positional data to achieve different ends." | |
| The new work by King's group contains three firsts: | |
|
|
|
| King points out that employing a 3D local observable opens exciting new avenues in multi-dimensional AFM in perturbative operating conditions. "One future application of particular interest may be in mapping the trajectory of flexible and disordered protein domains," he says. "There is a growing consensus within the biophysics community that this type of molecular motion is crucial – e.g., in boosting probability of binding and enhancing capture radii in protein-protein interactions – but it is very challenging to address via traditional techniques." | |
| He also emphasizes that the team's AFM experiments are performed in operating conditions that are common: tapping mode, commercial tips, in fluid, at room temperature; hence, they should be widely applicable across the many fields that routinely use AFMs. | |
 By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
By
Michael
Berger
– Michael is author of three books by the Royal Society of Chemistry:
Nano-Society: Pushing the Boundaries of Technology,
Nanotechnology: The Future is Tiny, and
Nanoengineering: The Skills and Tools Making Technology Invisible
Copyright ©
Nanowerk LLC
|
|
|
Become a Spotlight guest author! Join our large and growing group of guest contributors. Have you just published a scientific paper or have other exciting developments to share with the nanotechnology community? Here is how to publish on nanowerk.com. |
|
