Showing Spotlights 73 - 80 of 110 in category All (newest first):
 The folks at the European NanoHand project, whose nanogripper design we have covered in a previous Nanowerk Spotlight, seem to have loved playing with their plastic toy kits as kids. At least that's the impression you get when watching their latest video explaining their proof-of-principle study of scanning probe tips defined by planar nanolithography and integrated with AFM probes using nanomanipulation. They have prefabricated nanoscale needles, to be picked up by nanogrippers inside a scanning electron microscope. They then use these nanobits as ultralong tips in atomic force microscopes. The researchers call the needles 'nanobits' because they are a reminder of drill bits - you can have a library of different nanobits and then pick the one you want, and mount it where you want it.
The folks at the European NanoHand project, whose nanogripper design we have covered in a previous Nanowerk Spotlight, seem to have loved playing with their plastic toy kits as kids. At least that's the impression you get when watching their latest video explaining their proof-of-principle study of scanning probe tips defined by planar nanolithography and integrated with AFM probes using nanomanipulation. They have prefabricated nanoscale needles, to be picked up by nanogrippers inside a scanning electron microscope. They then use these nanobits as ultralong tips in atomic force microscopes. The researchers call the needles 'nanobits' because they are a reminder of drill bits - you can have a library of different nanobits and then pick the one you want, and mount it where you want it.
Sep 9th, 2009
 A Japanese-UK research team has now demonstrated that cryoEM image analysis may be exploited to obtain structural information of sufficient resolution to reveal the absolute three-dimensional (3D) configuration of a designed DNA nanostructure. With their technique they have obtained structural information at sufficient resolution to visualize the DNA helix and reveal the absolute stereochemistry of a self-assembled DNA tetrahedron. Each edge is a 7 nm, 20 base pair duplex, and the edges are connected covalently through single unpaired adenosine nucleotides, making it a rigid, triangulated structure that could serve as a building block for larger 3D structures or as a molecular cage. This DNA tetrahedron is the smallest 3D nanostructure made by DNA self-assembly.
A Japanese-UK research team has now demonstrated that cryoEM image analysis may be exploited to obtain structural information of sufficient resolution to reveal the absolute three-dimensional (3D) configuration of a designed DNA nanostructure. With their technique they have obtained structural information at sufficient resolution to visualize the DNA helix and reveal the absolute stereochemistry of a self-assembled DNA tetrahedron. Each edge is a 7 nm, 20 base pair duplex, and the edges are connected covalently through single unpaired adenosine nucleotides, making it a rigid, triangulated structure that could serve as a building block for larger 3D structures or as a molecular cage. This DNA tetrahedron is the smallest 3D nanostructure made by DNA self-assembly.
Aug 25th, 2009
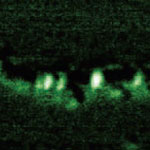 It its more than 25 years of existence, Scanning Tunneling Microscopy has predominantly brought us extremely detailed images of matter at the molecular and atomic level. The Scanning Tunneling Microscope (STM) is a non-optical microscope that scans an electrical probe over a surface to be imaged to detect a weak electric current flowing between the tip and the surface. It allows scientists to visualize regions of high electron density and hence infer the position of individual atoms and molecules on the surface of a lattice. Now, researchers in Japan have managed to partially sequence a single DNA molecule with a STM - a significant step towards the realization of electronic-based single-molecule DNA sequencing.
It its more than 25 years of existence, Scanning Tunneling Microscopy has predominantly brought us extremely detailed images of matter at the molecular and atomic level. The Scanning Tunneling Microscope (STM) is a non-optical microscope that scans an electrical probe over a surface to be imaged to detect a weak electric current flowing between the tip and the surface. It allows scientists to visualize regions of high electron density and hence infer the position of individual atoms and molecules on the surface of a lattice. Now, researchers in Japan have managed to partially sequence a single DNA molecule with a STM - a significant step towards the realization of electronic-based single-molecule DNA sequencing.
Aug 12th, 2009
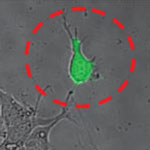 The Atomic Force Microscope (AFM) is a key tool for nanotechnology. This instrument has become the most widely used tool for imaging, measuring and manipulating matter at the nanoscale and in turn has inspired a variety of other scanning probe techniques. Originally the AFM was used to image the topography of surfaces, but by modifying the tip it is possible to measure other quantities (for example, electric and magnetic properties, chemical potentials, friction and so on), and also to perform various types of spectroscopy and analysis. Increasingly, the AFM is also becoming a tool for nanofabrication. Relatively new is the use of AFM in cell biology. Researchers in Switzerland have now demonstrated novel cell biology applications using hollow force-controlled AFM cantilevers - a new device they have called FluidFM.
The Atomic Force Microscope (AFM) is a key tool for nanotechnology. This instrument has become the most widely used tool for imaging, measuring and manipulating matter at the nanoscale and in turn has inspired a variety of other scanning probe techniques. Originally the AFM was used to image the topography of surfaces, but by modifying the tip it is possible to measure other quantities (for example, electric and magnetic properties, chemical potentials, friction and so on), and also to perform various types of spectroscopy and analysis. Increasingly, the AFM is also becoming a tool for nanofabrication. Relatively new is the use of AFM in cell biology. Researchers in Switzerland have now demonstrated novel cell biology applications using hollow force-controlled AFM cantilevers - a new device they have called FluidFM.
Jul 8th, 2009
 A major concern in microbiology is to determine whether a bacterium is dead or alive. This crucial question has major consequences in food industry, water supply or health care. While culture-based tests can determine whether bacteria can proliferate and form colonies, these tests are time-consuming and work poorly with certain slow-growing or non-culturable bacteria. They are not suitable for applications where real-time results are needed, e.g. in industrial manufacturing or food processing. A team of scientists in France has now discovered that living and dead cells can be discriminated with a nanotechnology technique on the basis of their cell wall nanomechanical properties.
A major concern in microbiology is to determine whether a bacterium is dead or alive. This crucial question has major consequences in food industry, water supply or health care. While culture-based tests can determine whether bacteria can proliferate and form colonies, these tests are time-consuming and work poorly with certain slow-growing or non-culturable bacteria. They are not suitable for applications where real-time results are needed, e.g. in industrial manufacturing or food processing. A team of scientists in France has now discovered that living and dead cells can be discriminated with a nanotechnology technique on the basis of their cell wall nanomechanical properties.
May 26th, 2009
 For the past 20+ years, the atomic force microscope has been one of the most important tools to visualize nanoscale objects where conventional optics cannot resolve them due to the wave nature of light. One limiting factor of conventional AFM operation is the speed at which images can be acquired. Over the past five years, researchers have been developing a high-speed AFM capable of video-rate image capture. An AFM with this ability enables nanoscale processes to be observed in real-time, rather than capturing only snap-shots in time. An obvious application of this instrument is to modify the sample surface while observing changes in the surface topography. Successful demonstration of this would indicate the potential for a new generation of fabrication tools. Scientists have now done exactly that.
For the past 20+ years, the atomic force microscope has been one of the most important tools to visualize nanoscale objects where conventional optics cannot resolve them due to the wave nature of light. One limiting factor of conventional AFM operation is the speed at which images can be acquired. Over the past five years, researchers have been developing a high-speed AFM capable of video-rate image capture. An AFM with this ability enables nanoscale processes to be observed in real-time, rather than capturing only snap-shots in time. An obvious application of this instrument is to modify the sample surface while observing changes in the surface topography. Successful demonstration of this would indicate the potential for a new generation of fabrication tools. Scientists have now done exactly that.
Mar 16th, 2009
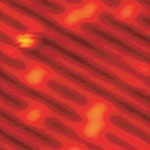 Over the past 25 years, Scanning Tunneling Microscopy (STM) has brought us extremely detailed images of matter at the molecular and atomic level. STM, which is a non-optical technique, works by scanning an electrical probe over a surface to be imaged to detect a weak electric current flowing between the tip and the surface. The STM allows scientists to visualize regions of high electron density and hence infer the position of individual atoms and molecules on the surface of a lattice.
Researchers also believe that the strength of time-lapsed high-resolution STM work to unravel complex surface reactions would allow them to achieve one of the 'Holy Grails' within the area of surface science, which is to directly observe chemical reactions at the atomic scale. A research team in Denmark has now shown that, by means of high-resolution STM studies in conjunction with density functional theory calculations, it is possible to follow the intermediate steps of a complex oxidation reaction.
Over the past 25 years, Scanning Tunneling Microscopy (STM) has brought us extremely detailed images of matter at the molecular and atomic level. STM, which is a non-optical technique, works by scanning an electrical probe over a surface to be imaged to detect a weak electric current flowing between the tip and the surface. The STM allows scientists to visualize regions of high electron density and hence infer the position of individual atoms and molecules on the surface of a lattice.
Researchers also believe that the strength of time-lapsed high-resolution STM work to unravel complex surface reactions would allow them to achieve one of the 'Holy Grails' within the area of surface science, which is to directly observe chemical reactions at the atomic scale. A research team in Denmark has now shown that, by means of high-resolution STM studies in conjunction with density functional theory calculations, it is possible to follow the intermediate steps of a complex oxidation reaction.
Mar 3rd, 2009
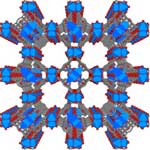 Crystalline nanoporous compounds have attracted the attention of scientists and materials engineers because of the interest in creating nanoscale spaces and the novel phenomena in them. Nanoporous materials find applications in many chemical processes such as separation and catalysis and are also heavily researched as storage materials, for instance in hydrogen fuel cells. Researchers still lack a complete understanding of the mechanism that leads to, and occurs during, crystal growth. Only once scientists achieve full control of properties such as composition, structure, size, morphology, and the presence and form of defects within the crystals can they fully exploit the benefits of crystalline nanoporous materials for the fabrication of novel materials. Researchers in the UK have now presented definitive real-time evidence of the crystal growth mechanism in what appears to be the first high-resolution observation of in-situ crystal growth of a crystalline nanoporous material monitored using atomic force microscopy (AFM).
Crystalline nanoporous compounds have attracted the attention of scientists and materials engineers because of the interest in creating nanoscale spaces and the novel phenomena in them. Nanoporous materials find applications in many chemical processes such as separation and catalysis and are also heavily researched as storage materials, for instance in hydrogen fuel cells. Researchers still lack a complete understanding of the mechanism that leads to, and occurs during, crystal growth. Only once scientists achieve full control of properties such as composition, structure, size, morphology, and the presence and form of defects within the crystals can they fully exploit the benefits of crystalline nanoporous materials for the fabrication of novel materials. Researchers in the UK have now presented definitive real-time evidence of the crystal growth mechanism in what appears to be the first high-resolution observation of in-situ crystal growth of a crystalline nanoporous material monitored using atomic force microscopy (AFM).
Oct 30th, 2008
 The folks at the European NanoHand project, whose nanogripper design we have covered in a previous Nanowerk Spotlight, seem to have loved playing with their plastic toy kits as kids. At least that's the impression you get when watching their latest video explaining their proof-of-principle study of scanning probe tips defined by planar nanolithography and integrated with AFM probes using nanomanipulation. They have prefabricated nanoscale needles, to be picked up by nanogrippers inside a scanning electron microscope. They then use these nanobits as ultralong tips in atomic force microscopes. The researchers call the needles 'nanobits' because they are a reminder of drill bits - you can have a library of different nanobits and then pick the one you want, and mount it where you want it.
The folks at the European NanoHand project, whose nanogripper design we have covered in a previous Nanowerk Spotlight, seem to have loved playing with their plastic toy kits as kids. At least that's the impression you get when watching their latest video explaining their proof-of-principle study of scanning probe tips defined by planar nanolithography and integrated with AFM probes using nanomanipulation. They have prefabricated nanoscale needles, to be picked up by nanogrippers inside a scanning electron microscope. They then use these nanobits as ultralong tips in atomic force microscopes. The researchers call the needles 'nanobits' because they are a reminder of drill bits - you can have a library of different nanobits and then pick the one you want, and mount it where you want it.
 Subscribe to our Nanotechnology Spotlight feed
Subscribe to our Nanotechnology Spotlight feed





